Epidermal growth factor receptor
| VALUE_ERROR (nil) | |||||||
|---|---|---|---|---|---|---|---|
| Identifiers | |||||||
| Aliases | |||||||
| External IDs | GeneCards: [1] | ||||||
| Orthologs | |||||||
| Species | Human | Mouse | |||||
| Entrez |
|
| |||||
| Ensembl |
|
| |||||
| UniProt |
|
| |||||
| RefSeq (mRNA) |
|
| |||||
| RefSeq (protein) |
|
| |||||
| Location (UCSC) | n/a | n/a | |||||
| PubMed search | n/a | n/a | |||||
| Wikidata | |||||||
| |||||||
The epidermal growth factor receptor (EGFR; ErbB-1; HER1 in humans) is a transmembrane protein that is a receptor for members of the epidermal growth factor family (EGF family) of extracellular protein ligands.[1]
The epidermal growth factor receptor is a member of the ErbB family of receptors, a subfamily of four closely related receptor tyrosine kinases: EGFR (ErbB-1), HER2/neu (ErbB-2), Her 3 (ErbB-3) and Her 4 (ErbB-4). In many cancer types, mutations affecting EGFR expression or activity could result in cancer.[2]
Epidermal growth factor and its receptor was discovered by Stanley Cohen of Vanderbilt University. Cohen shared the 1986 Nobel Prize in Medicine with Rita Levi-Montalcini for their discovery of growth factors.
Deficient signaling of the EGFR and other receptor tyrosine kinases in humans is associated with diseases such as Alzheimer's, while over-expression is associated with the development of a wide variety of tumors. Interruption of EGFR signalling, either by blocking EGFR binding sites on the extracellular domain of the receptor or by inhibiting intracellular tyrosine kinase activity, can prevent the growth of EGFR-expressing tumours and improve the patient's condition.
Function
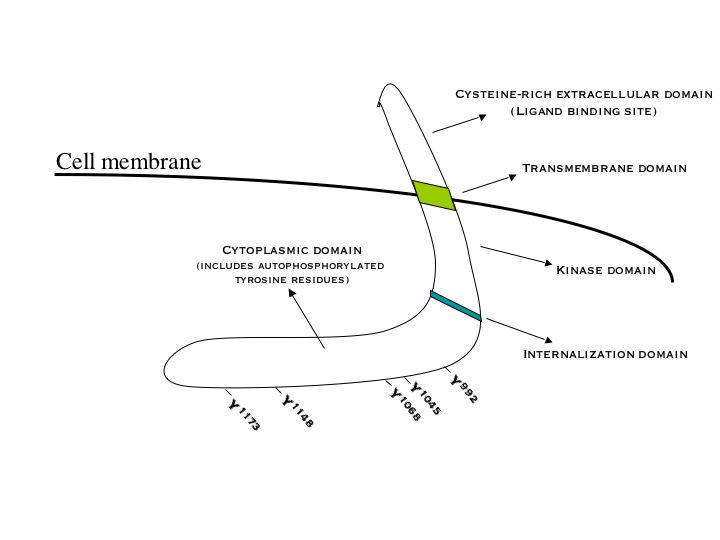
Epidermal growth factor receptor (EGFR) is a transmembrane protein that is activated by binding of its specific ligands, including epidermal growth factor and transforming growth factor α (TGFα)[3] ErbB2 has no known direct activating ligand, and may be in an activated state constitutively or become active upon heterodimerization with other family members such as EGFR. Upon activation by its growth factor ligands, EGFR undergoes a transition from an inactive monomeric form to an active homodimer.[4] – although there is some evidence that preformed inactive dimers may also exist before ligand binding.[citation needed] In addition to forming homodimers after ligand binding, EGFR may pair with another member of the ErbB receptor family, such as ErbB2/Her2/neu, to create an activated heterodimer. There is also evidence to suggest that clusters of activated EGFRs form, although it remains unclear whether this clustering is important for activation itself or occurs subsequent to activation of individual dimers.[citation needed]
EGFR dimerization stimulates its intrinsic intracellular protein-tyrosine kinase activity. As a result, autophosphorylation of several tyrosine (Y) residues in the C-terminal domain of EGFR occurs. These include Y992, Y1045, Y1068, Y1148 and Y1173, as shown in the adjacent diagram.[5] This autophosphorylation elicits downstream activation and signaling by several other proteins that associate with the phosphorylated tyrosines through their own phosphotyrosine-binding SH2 domains. These downstream signaling proteins initiate several signal transduction cascades, principally the MAPK, Akt and JNK pathways, leading to DNA synthesis and cell proliferation.[6] Such proteins modulate phenotypes such as cell migration, adhesion, and proliferation. Activation of the receptor is important for the innate immune response in human skin. The kinase domain of EGFR can also cross-phosphorylate tyrosine residues of other receptors it is aggregated with, and can itself be activated in that manner.
Biological roles
The EGFR is essential for ductal development of the mammary glands,[7][8][9] and agonists of the EGFR such as amphiregulin, TGF-α, and heregulin induce both ductal and lobuloalveolar development even in the absence of estrogen and progesterone.[10][11]
Role in human disease
Cancer
Mutations that lead to EGFR overexpression (known as upregulation or amplification) have been associated with a number of cancers, including adenocarcinoma of the lung (40% of cases), anal cancers,[12] glioblastoma (50%) and epithelian tumors of the head and neck (80-100%).[13] These somatic mutations involving EGFR lead to its constant activation, which produces uncontrolled cell division.[14] In glioblastoma a specific mutation of EGFR, called EGFRvIII, is often observed.[15] Mutations, amplifications or misregulations of EGFR or family members are implicated in about 30% of all epithelial cancers.[citation needed]
Inflammatory disease
Aberrant EGFR signaling has been implicated in psoriasis, eczema and atherosclerosis.[16][17] However, its exact roles in these conditions are ill-defined.
Monogenic disease
A single child displaying multi-organ epithelial inflammation was found to have a homozygous loss of function mutation in the EGFR gene. The pathogenicity of the EGFR mutation was supported by in vitro experiments and functional analysis of a skin biopsy. His severe phenotype reflects many previous research findings into EGFR function. His clinical features included a papulopustular rash, dry skin, chronic diarrhoea, abnormalities of hair growth, breathing difficulties and electrolyte imbalances.[18]
Wound healing and fibrosis
EGFR has been shown to play a critical role in TGF-beta1 dependent fibroblast to myofibroblast differentiation.[19][20] Aberrant persistence of myofibroblasts within tissues can lead to progressive tissue fibrosis, impairing tissue or organ function (e.g. skin hypertrophic or keloid scars, liver cirrhosis, myocardial fibrosis, chronic kidney disease).
Medical applications
Drug target
The identification of EGFR as an oncogene has led to the development of anticancer therapeutics directed against EGFR (called "EGFR inhibitors"), including gefitinib,[21] erlotinib, afatinib, brigatinib and icotinib[22] for lung cancer, and cetuximab for colon cancer. More recently AstraZeneca has developed Osimertinib, a third generation tyrosine kinase inhibitor.[23]
Many therapeutic approaches are aimed at the EGFR. Cetuximab and panitumumab are examples of monoclonal antibody inhibitors. However the former is of the IgG1 type, the latter of the IgG2 type; consequences on antibody-dependent cellular cytotoxicity can be quite different.[24] Other monoclonals in clinical development are zalutumumab, nimotuzumab, and matuzumab. The monoclonal antibodies block the extracellular ligand binding domain. With the binding site blocked, signal molecules can no longer attach there and activate the tyrosine kinase.
Another method is using small molecules to inhibit the EGFR tyrosine kinase, which is on the cytoplasmic side of the receptor. Without kinase activity, EGFR is unable to activate itself, which is a prerequisite for binding of downstream adaptor proteins. Ostensibly by halting the signaling cascade in cells that rely on this pathway for growth, tumor proliferation and migration is diminished. Gefitinib, erlotinib, brigatinib and lapatinib (mixed EGFR and ERBB2 inhibitor) are examples of small molecule kinase inhibitors.
CimaVax-EGF, an active vaccine targeting EGF as the major ligand of EGF, uses a different approach, raising antibodies against EGF itself, thereby denying EGFR-dependent cancers of a proliferative stimulus;[25] it is in use as a cancer therapy against non-small-cell lung carcinoma (the most common form of lung cancer) in Cuba, and is undergoing further trials for possible licensing in Japan, Europe, and the United States.[26]
There are several quantitative methods available that use protein phosphorylation detection to identify EGFR family inhibitors.[27]
New drugs such as osimertinib, gefitinib, erlotinib and brigatinib directly target the EGFR. Patients have been divided into EGFR-positive and EGFR-negative, based upon whether a tissue test shows a mutation. EGFR-positive patients have shown a 60% response rate, which exceeds the response rate for conventional chemotherapy.[28]
However, many patients develop resistance. Two primary sources of resistance are the T790M Mutation and MET oncogene.[28] However, as of 2010 there was no consensus of an accepted approach to combat resistance nor FDA approval of a specific combination. Clinical trial phase II results reported for brigatinib targeting the T790M mutation, and brigatinib received Breakthrough Therapy designation status by FDA in Feb. 2015.
The most common adverse effect of EGFR inhibitors, found in more than 90% of patients, is a papulopustular rash that spreads across the face and torso; the rash's presence is correlated with the drug's antitumor effect.[29] In 10% to 15% of patients the effects can be serious and require treatment.[30][31]
Some tests are aiming at predicting benefit from EGFR treatment, as Veristrat.[32]
Laboratory research using genetically engineered stem cells to target EGFR in mice was reported in 2014 to show promise.[33] EGFR is a well-established target for monoclonal antibodies and specific tyrosine kinase inhibitors.[34]
Target for imaging agents
Imaging agents have been developed which identify EGFR-dependent cancers using labeled EGF.[35] The feasibility of in vivo imaging of EGFR expression has been demonstrated in several studies.[36][37]
Interactions
Epidermal growth factor receptor has been shown to interact with:
- AR,[38][39]
- ARF4,[40]
- CAV1,[41]
- CAV3,[41]
- CBL,[42][43][44][45][46]
- CBLB,[43][47]
- CBLC,[48][49]
- CD44,[19]
- CDC25A,[50]
- CRK,[47][51]
- CTNNB1,[52][53][54]
- DCN,[55][56]
- EGF,[57][58]
- GRB14,[59]
- Grb2,[47][57][59][60][61][62][63][64][65][66]
- JAK2,[67]
- MUC1,[68][69]
- NCK1,[60][70][71]
- NCK2[60][72][73]
- PKC alpha,[74]
- PLCG1,[42][75]
- PLSCR1,[76]
- PTPN1,[77][78]
- PTPN11,[47][79]
- PTPN6,[79][80]
- PTPRK,[81]
- SH2D3A,[82]
- SH3KBP1,[83][84]
- SHC1,[47][85]
- SOS1,[65][86][87]
- Src,[67][88][89]
- STAT1,[67][90]
- STAT3,[67][91]
- STAT5A,[47][67]
- UBC,[44][45][92] and
- WAS.[93]
In fruitflies, the epidermal growth factor receptor interacts with Spitz.[94]
References
- ↑ Herbst RS (2004). "Review of epidermal growth factor receptor biology". International Journal of Radiation Oncology, Biology, Physics. 59 (2 Suppl): 21–6. doi:10.1016/j.ijrobp.2003.11.041. PMID 15142631.
- ↑ Zhang H, Berezov A, Wang Q, Zhang G, Drebin J, Murali R, Greene MI (August 2007). "ErbB receptors: from oncogenes to targeted cancer treatment". The Journal of Clinical Investigation. 117 (8): 2051–8. doi:10.1172/JCI32278. PMC 1934579. PMID 17671639.
- ↑ note, a full list of the ligands able to activate EGFR and other members of the ErbB family is given in the ErbB article).
- ↑ Yarden Y, Schlessinger J (March 1987). "Epidermal growth factor induces rapid, reversible aggregation of the purified epidermal growth factor receptor". Biochemistry. 26 (5): 1443–51. doi:10.1021/bi00379a035. PMID 3494473.
- ↑ Downward J, Parker P, Waterfield MD (1984). "Autophosphorylation sites on the epidermal growth factor receptor". Nature. 311 (5985): 483–5. doi:10.1038/311483a0. PMID 6090945.
- ↑ Oda K, Matsuoka Y, Funahashi A, Kitano H (2005). "A comprehensive pathway map of epidermal growth factor receptor signaling". Molecular Systems Biology. 1 (1): 2005.0010. doi:10.1038/msb4100014. PMC 1681468. PMID 16729045.
- ↑ Sebastian J, Richards RG, Walker MP, Wiesen JF, Werb Z, Derynck R, Hom YK, Cunha GR, DiAugustine RP (September 1998). "Activation and function of the epidermal growth factor receptor and erbB-2 during mammary gland morphogenesis". Cell Growth & Differentiation. 9 (9): 777–85. PMID 9751121.
- ↑ McBryan J, Howlin J, Napoletano S, Martin F (June 2008). "Amphiregulin: role in mammary gland development and breast cancer". Journal of Mammary Gland Biology and Neoplasia. 13 (2): 159–69. doi:10.1007/s10911-008-9075-7. PMID 18398673.
- ↑ Sternlicht MD, Sunnarborg SW (June 2008). "The ADAM17-amphiregulin-EGFR axis in mammary development and cancer". Journal of Mammary Gland Biology and Neoplasia. 13 (2): 181–94. doi:10.1007/s10911-008-9084-6. PMC 2723838. PMID 18470483.
- ↑ Kenney NJ, Bowman A, Korach KS, Barrett JC, Salomon DS (May 2003). "Effect of exogenous epidermal-like growth factors on mammary gland development and differentiation in the estrogen receptor-alpha knockout (ERKO) mouse". Breast Cancer Research and Treatment. 79 (2): 161–73. doi:10.1023/a:1023938510508. PMID 12825851.
- ↑ Kenney NJ, Smith GH, Rosenberg K, Cutler ML, Dickson RB (December 1996). "Induction of ductal morphogenesis and lobular hyperplasia by amphiregulin in the mouse mammary gland". Cell Growth & Differentiation. 7 (12): 1769–81. PMID 8959346.
- ↑ Walker F, Abramowitz L, Benabderrahmane D, Duval X, Descatoire V, Hénin D, Lehy T, Aparicio T (November 2009). "Growth factor receptor expression in anal squamous lesions: modifications associated with oncogenic human papillomavirus and human immunodeficiency virus". Human Pathology. 40 (11): 1517–27. doi:10.1016/j.humpath.2009.05.010. PMID 19716155.
- ↑ Kumar V, Abbas A, Aster J (2013). Robbins basic pathology. Philadelphia: Elsevier/Saunders. p. 179. ISBN 9781437717815.
- ↑ Lynch TJ, Bell DW, Sordella R, Gurubhagavatula S, Okimoto RA, Brannigan BW, Harris PL, Haserlat SM, Supko JG, Haluska FG, Louis DN, Christiani DC, Settleman J, Haber DA (May 2004). "Activating mutations in the epidermal growth factor receptor underlying responsiveness of non-small-cell lung cancer to gefitinib". The New England Journal of Medicine. 350 (21): 2129–39. doi:10.1056/NEJMoa040938. PMID 15118073.
- ↑ Kuan CT, Wikstrand CJ, Bigner DD (June 2001). "EGF mutant receptor vIII as a molecular target in cancer therapy". Endocrine-Related Cancer. 8 (2): 83–96. doi:10.1677/erc.0.0080083. PMID 11397666.
- ↑ Jost M, Kari C, Rodeck U (2000). "The EGF receptor - an essential regulator of multiple epidermal functions". European Journal of Dermatology. 10 (7): 505–10. PMID 11056418.
- ↑ Dreux AC, Lamb DJ, Modjtahedi H, Ferns GA (May 2006). "The epidermal growth factor receptors and their family of ligands: their putative role in atherogenesis". Atherosclerosis. 186 (1): 38–53. doi:10.1016/j.atherosclerosis.2005.06.038. PMID 16076471.
- ↑ Campbell P, Morton PE, Takeichi T, Salam A, Roberts N, Proudfoot LE, Mellerio JE, Aminu K, Wellington C, Patil SN, Akiyama M, Liu L, McMillan JR, Aristodemou S, Ishida-Yamamoto A, Abdul-Wahab A, Petrof G, Fong K, Harnchoowong S, Stone KL, Harper JI, McLean WH, Simpson MA, Parsons M, McGrath JA (October 2014). "Epithelial inflammation resulting from an inherited loss-of-function mutation in EGFR". The Journal of Investigative Dermatology. 134 (10): 2570–8. doi:10.1038/jid.2014.164. PMC 4090136. PMID 24691054.
- ↑ 19.0 19.1 Midgley AC, Rogers M, Hallett MB, Clayton A, Bowen T, Phillips AO, Steadman R (May 2013). "Transforming growth factor-β1 (TGF-β1)-stimulated fibroblast to myofibroblast differentiation is mediated by hyaluronan (HA)-facilitated epidermal growth factor receptor (EGFR) and CD44 co-localization in lipid rafts". The Journal of Biological Chemistry. 288 (21): 14824–38. doi:10.1074/jbc.M113.451336. PMC 3663506. PMID 23589287.
- ↑ Midgley AC, Bowen T, Phillips AO, Steadman R (April 2014). "MicroRNA-7 inhibition rescues age-associated loss of epidermal growth factor receptor and hyaluronan-dependent differentiation in fibroblasts". Aging Cell. 13 (2): 235–44. doi:10.1111/acel.12167. PMC 4331777. PMID 24134702.
- ↑ Paez JG, Jänne PA, Lee JC, Tracy S, Greulich H, Gabriel S, Herman P, Kaye FJ, Lindeman N, Boggon TJ, Naoki K, Sasaki H, Fujii Y, Eck MJ, Sellers WR, Johnson BE, Meyerson M (June 2004). "EGFR mutations in lung cancer: correlation with clinical response to gefitinib therapy". Science. 304 (5676): 1497–500. doi:10.1126/science.1099314. PMID 15118125.
- ↑ Liang W, Wu X, Fang W, Zhao Y, Yang Y, Hu Z, Xue C, Zhang J, Zhang J, Ma Y, Zhou T, Yan Y, Hou X, Qin T, Dinglin X, Tian Y, Huang P, Huang Y, Zhao H, Zhang L (12 February 2014). "Network meta-analysis of erlotinib, gefitinib, afatinib and icotinib in patients with advanced non-small-cell lung cancer harboring EGFR mutations". PLoS ONE. 9 (2): e85245. doi:10.1371/journal.pone.0085245. PMC 3922700. PMID 24533047.
- ↑ Greig SL (February 2016). "Osimertinib: First Global Approval". Drugs. 76 (2): 263–73. doi:10.1007/s40265-015-0533-4. PMID 26729184.
- ↑ Yan L, Beckman RA (October 2005). "Pharmacogenetics and pharmacogenomics in oncology therapeutic antibody development". BioTechniques. 39 (4): 565–8. doi:10.2144/000112043. PMID 16235569.
- ↑ Rodríguez PC, Rodríguez G, González G, Lage A (Winter 2010). "Clinical development and perspectives of CIMAvax EGF, Cuban vaccine for non-small-cell lung cancer therapy". MEDICC Review. 12 (1): 17–23. PMID 20387330.
- ↑ Patel N (11 May 2015). "Cuba Has a Lung Cancer Vaccine—And America Wants It". Wired. Retrieved 13 May 2015.
- ↑ Olive DM (October 2004). "Quantitative methods for the analysis of protein phosphorylation in drug development". Expert Review of Proteomics. 1 (3): 327–41. doi:10.1586/14789450.1.3.327. PMID 15966829.
- ↑ 28.0 28.1 Jackman DM, Miller VA, Cioffredi LA, Yeap BY, Jänne PA, Riely GJ, Ruiz MG, Giaccone G, Sequist LV, Johnson BE (August 2009). "Impact of epidermal growth factor receptor and KRAS mutations on clinical outcomes in previously untreated non-small cell lung cancer patients: results of an online tumor registry of clinical trials". Clinical Cancer Research. 15 (16): 5267–73. doi:10.1158/1078-0432.CCR-09-0888. PMC 3219530. PMID 19671843.
- ↑ Liu HB, Wu Y, Lv TF, Yao YW, Xiao YY, Yuan DM, Song Y (2013). "Skin rash could predict the response to EGFR tyrosine kinase inhibitor and the prognosis for patients with non-small cell lung cancer: a systematic review and meta-analysis". PLoS ONE. 8 (1): e55128. doi:10.1371/journal.pone.0055128. PMC 3559430. PMID 23383079.
- ↑ Gerber PA, Meller S, Eames T, Buhren BA, Schrumpf H, Hetzer S, Ehmann LM, Budach W, Bölke E, Matuschek C, Wollenberg A, Homey B (2012). "Management of EGFR-inhibitor associated rash: a retrospective study in 49 patients". European Journal of Medical Research. 17 (1): 4. doi:10.1186/2047-783X-17-4. PMC 3351712. PMID 22472354.
- ↑ Lacouture ME (October 2006). "Mechanisms of cutaneous toxicities to EGFR inhibitors". Nature Reviews. Cancer. 6 (10): 803–12. doi:10.1038/nrc1970. PMID 16990857.
- ↑ Molina-Pinelo S, Pastor MD, Paz-Ares L (February 2014). "VeriStrat: a prognostic and/or predictive biomarker for advanced lung cancer patients?". Expert Review of Respiratory Medicine. 8 (1): 1–4. doi:10.1586/17476348.2014.861744. PMID 24308656.
- ↑ Stuckey DW, Hingtgen SD, Karakas N, Rich BE, Shah K (February 2015). "Engineering toxin-resistant therapeutic stem cells to treat brain tumors". Stem Cells. 33 (2): 589–600. doi:10.1002/stem.1874. PMC 4305025. PMID 25346520.
- ↑ Roskoski R Jr (2014). "The ErbB/HER family of protein-tyrosine kinases and cancer". Pharmacol Res. 79: 34–74. doi:10.1016/j.phrs.2013.11.002. PMID 24269963.
- ↑ Lucas LJ, Tellez CA, Castilho ML, Lee CL, Hupman MA, Vieira LS, Ferreira I, Raniero L, Hewitt KC (May 2015). "Development of a sensitive, stable and EGFR-specific molecular imaging agent for surface enhanced Raman spectroscopy". Journal of Raman Spectroscopy. 46 (5): 434–446. doi:10.1002/jrs.4678.
- ↑ Lucas LJ, Chen XK, Smith AJ, Korbelik M, Zeng, Haitian L, Lee PW, Hewitt KC (23 January 2015). "Aggregation of nanoparticles in endosomes and lysosomes produces surface-enhanced Raman spectroscopy". Journal of Nanophotonics. 9 (1): 093094–1–14. doi:10.1117/1.JNP.9.093094.
- ↑ Andersson KG, Oroujeni M, Garousi J, Mitran B, Ståhl S, Orlova A, Löfblom J, Tolmachev V (December 2016). "Feasibility of imaging of epidermal growth factor receptor expression with ZEGFR:2377 affibody molecule labeled with 99mTc using a peptide-based cysteine-containing chelator". International Journal of Oncology. 49 (6): 2285–2293. doi:10.3892/ijo.2016.3721. PMC 5118000. PMID 27748899.
- ↑ Bonaccorsi L, Carloni V, Muratori M, Formigli L, Zecchi S, Forti G, Baldi E (October 2004). "EGF receptor (EGFR) signaling promoting invasion is disrupted in androgen-sensitive prostate cancer cells by an interaction between EGFR and androgen receptor (AR)". International Journal of Cancer. 112 (1): 78–86. doi:10.1002/ijc.20362. PMID 15305378.
- ↑ Bonaccorsi L, Muratori M, Carloni V, Marchiani S, Formigli L, Forti G, Baldi E (August 2004). "The androgen receptor associates with the epidermal growth factor receptor in androgen-sensitive prostate cancer cells". Steroids. 69 (8–9): 549–52. doi:10.1016/j.steroids.2004.05.011. PMID 15288768.
- ↑ Kim SW, Hayashi M, Lo JF, Yang Y, Yoo JS, Lee JD (January 2003). "ADP-ribosylation factor 4 small GTPase mediates epidermal growth factor receptor-dependent phospholipase D2 activation". The Journal of Biological Chemistry. 278 (4): 2661–8. doi:10.1074/jbc.M205819200. PMID 12446727.
- ↑ 41.0 41.1 Couet J, Sargiacomo M, Lisanti MP (November 1997). "Interaction of a receptor tyrosine kinase, EGF-R, with caveolins. Caveolin binding negatively regulates tyrosine and serine/threonine kinase activities". The Journal of Biological Chemistry. 272 (48): 30429–38. doi:10.1074/jbc.272.48.30429. PMID 9374534.
- ↑ 42.0 42.1 Tvorogov D, Carpenter G (July 2002). "EGF-dependent association of phospholipase C-gamma1 with c-Cbl". Experimental Cell Research. 277 (1): 86–94. doi:10.1006/excr.2002.5545. PMID 12061819.
- ↑ 43.0 43.1 Ettenberg SA, Keane MM, Nau MM, Frankel M, Wang LM, Pierce JH, Lipkowitz S (March 1999). "cbl-b inhibits epidermal growth factor receptor signaling". Oncogene. 18 (10): 1855–66. doi:10.1038/sj.onc.1202499. PMID 10086340.
- ↑ 44.0 44.1 Pennock S, Wang Z (May 2008). "A tale of two Cbls: interplay of c-Cbl and Cbl-b in epidermal growth factor receptor downregulation". Molecular and Cellular Biology. 28 (9): 3020–37. doi:10.1128/MCB.01809-07. PMC 2293090. PMID 18316398.
- ↑ 45.0 45.1 Umebayashi K, Stenmark H, Yoshimori T (August 2008). "Ubc4/5 and c-Cbl continue to ubiquitinate EGF receptor after internalization to facilitate polyubiquitination and degradation". Molecular Biology of the Cell. 19 (8): 3454–62. doi:10.1091/mbc.E07-10-0988. PMC 2488299. PMID 18508924.
- ↑ Ng C, Jackson RA, Buschdorf JP, Sun Q, Guy GR, Sivaraman J (March 2008). "Structural basis for a novel intrapeptidyl H-bond and reverse binding of c-Cbl-TKB domain substrates". The EMBO Journal. 27 (5): 804–16. doi:10.1038/emboj.2008.18. PMC 2265755. PMID 18273061.
- ↑ 47.0 47.1 47.2 47.3 47.4 47.5 Schulze WX, Deng L, Mann M (2005). "Phosphotyrosine interactome of the ErbB-receptor kinase family". Molecular Systems Biology. 1 (1): 2005.0008. doi:10.1038/msb4100012. PMC 1681463. PMID 16729043.
- ↑ Kim M, Tezuka T, Suziki Y, Sugano S, Hirai M, Yamamoto T (October 1999). "Molecular cloning and characterization of a novel cbl-family gene, cbl-c". Gene. 239 (1): 145–54. doi:10.1016/S0378-1119(99)00356-X. PMID 10571044.
- ↑ Keane MM, Ettenberg SA, Nau MM, Banerjee P, Cuello M, Penninger J, Lipkowitz S (June 1999). "cbl-3: a new mammalian cbl family protein". Oncogene. 18 (22): 3365–75. doi:10.1038/sj.onc.1202753. PMID 10362357.
- ↑ Wang Z, Wang M, Lazo JS, Carr BI (May 2002). "Identification of epidermal growth factor receptor as a target of Cdc25A protein phosphatase". The Journal of Biological Chemistry. 277 (22): 19470–5. doi:10.1074/jbc.M201097200. PMID 11912208.
- ↑ Hashimoto Y, Katayama H, Kiyokawa E, Ota S, Kurata T, Gotoh N, Otsuka N, Shibata M, Matsuda M (July 1998). "Phosphorylation of CrkII adaptor protein at tyrosine 221 by epidermal growth factor receptor". The Journal of Biological Chemistry. 273 (27): 17186–91. doi:10.1074/jbc.273.27.17186. PMID 9642287.
- ↑ Hazan RB, Norton L (April 1998). "The epidermal growth factor receptor modulates the interaction of E-cadherin with the actin cytoskeleton". The Journal of Biological Chemistry. 273 (15): 9078–84. doi:10.1074/jbc.273.15.9078. PMID 9535896.
- ↑ Schroeder JA, Adriance MC, McConnell EJ, Thompson MC, Pockaj B, Gendler SJ (June 2002). "ErbB-beta-catenin complexes are associated with human infiltrating ductal breast and murine mammary tumor virus (MMTV)-Wnt-1 and MMTV-c-Neu transgenic carcinomas". The Journal of Biological Chemistry. 277 (25): 22692–8. doi:10.1074/jbc.M201975200. PMID 11950845.
- ↑ Takahashi K, Suzuki K, Tsukatani Y (July 1997). "Induction of tyrosine phosphorylation and association of beta-catenin with EGF receptor upon tryptic digestion of quiescent cells at confluence". Oncogene. 15 (1): 71–8. doi:10.1038/sj.onc.1201160. PMID 9233779.
- ↑ Santra M, Reed CC, Iozzo RV (September 2002). "Decorin binds to a narrow region of the epidermal growth factor (EGF) receptor, partially overlapping but distinct from the EGF-binding epitope". The Journal of Biological Chemistry. 277 (38): 35671–81. doi:10.1074/jbc.M205317200. PMID 12105206.
- ↑ Iozzo RV, Moscatello DK, McQuillan DJ, Eichstetter I (February 1999). "Decorin is a biological ligand for the epidermal growth factor receptor". The Journal of Biological Chemistry. 274 (8): 4489–92. doi:10.1074/jbc.274.8.4489. PMID 9988678.
- ↑ 57.0 57.1 Wong L, Deb TB, Thompson SA, Wells A, Johnson GR (March 1999). "A differential requirement for the COOH-terminal region of the epidermal growth factor (EGF) receptor in amphiregulin and EGF mitogenic signaling". The Journal of Biological Chemistry. 274 (13): 8900–9. doi:10.1074/jbc.274.13.8900. PMID 10085134.
- ↑ Stortelers C, Souriau C, van Liempt E, van de Poll ML, van Zoelen EJ (July 2002). "Role of the N-terminus of epidermal growth factor in ErbB-2/ErbB-3 binding studied by phage display". Biochemistry. 41 (27): 8732–41. doi:10.1021/bi025878c. PMID 12093292.
- ↑ 59.0 59.1 Daly RJ, Sanderson GM, Janes PW, Sutherland RL (May 1996). "Cloning and characterization of GRB14, a novel member of the GRB7 gene family". The Journal of Biological Chemistry. 271 (21): 12502–10. doi:10.1074/jbc.271.21.12502. PMID 8647858.
- ↑ 60.0 60.1 60.2 Braverman LE, Quilliam LA (February 1999). "Identification of Grb4/Nckbeta, a src homology 2 and 3 domain-containing adapter protein having similar binding and biological properties to Nck". The Journal of Biological Chemistry. 274 (9): 5542–9. doi:10.1074/jbc.274.9.5542. PMID 10026169.
- ↑ Blagoev B, Kratchmarova I, Ong SE, Nielsen M, Foster LJ, Mann M (March 2003). "A proteomics strategy to elucidate functional protein-protein interactions applied to EGF signaling". Nature Biotechnology. 21 (3): 315–8. doi:10.1038/nbt790. PMID 12577067.
- ↑ Oneyama C, Nakano H, Sharma SV (March 2002). "UCS15A, a novel small molecule, SH3 domain-mediated protein-protein interaction blocking drug". Oncogene. 21 (13): 2037–50. doi:10.1038/sj.onc.1205271. PMID 11960376.
- ↑ Okutani T, Okabayashi Y, Kido Y, Sugimoto Y, Sakaguchi K, Matuoka K, Takenawa T, Kasuga M (December 1994). "Grb2/Ash binds directly to tyrosines 1068 and 1086 and indirectly to tyrosine 1148 of activated human epidermal growth factor receptors in intact cells". The Journal of Biological Chemistry. 269 (49): 31310–4. PMID 7527043.
- ↑ Tortora G, Damiano V, Bianco C, Baldassarre G, Bianco AR, Lanfrancone L, Pelicci PG, Ciardiello F (February 1997). "The RIalpha subunit of protein kinase A (PKA) binds to Grb2 and allows PKA interaction with the activated EGF-receptor". Oncogene. 14 (8): 923–8. doi:10.1038/sj.onc.1200906. PMID 9050991.
- ↑ 65.0 65.1 Buday L, Egan SE, Rodriguez Viciana P, Cantrell DA, Downward J (March 1994). "A complex of Grb2 adaptor protein, Sos exchange factor, and a 36-kDa membrane-bound tyrosine phosphoprotein is implicated in ras activation in T cells". The Journal of Biological Chemistry. 269 (12): 9019–23. PMID 7510700.
- ↑ Lowenstein EJ, Daly RJ, Batzer AG, Li W, Margolis B, Lammers R, Ullrich A, Skolnik EY, Bar-Sagi D, Schlessinger J (August 1992). "The SH2 and SH3 domain-containing protein GRB2 links receptor tyrosine kinases to ras signaling". Cell. 70 (3): 431–42. doi:10.1016/0092-8674(92)90167-B. PMID 1322798.
- ↑ 67.0 67.1 67.2 67.3 67.4 Olayioye MA, Beuvink I, Horsch K, Daly JM, Hynes NE (June 1999). "ErbB receptor-induced activation of stat transcription factors is mediated by Src tyrosine kinases". The Journal of Biological Chemistry. 274 (24): 17209–18. doi:10.1074/jbc.274.24.17209. PMID 10358079.
- ↑ Schroeder JA, Thompson MC, Gardner MM, Gendler SJ (April 2001). "Transgenic MUC1 interacts with epidermal growth factor receptor and correlates with mitogen-activated protein kinase activation in the mouse mammary gland". The Journal of Biological Chemistry. 276 (16): 13057–64. doi:10.1074/jbc.M011248200. PMID 11278868.
- ↑ Li Y, Ren J, Yu W, Li Q, Kuwahara H, Yin L, Carraway KL, Kufe D (September 2001). "The epidermal growth factor receptor regulates interaction of the human DF3/MUC1 carcinoma antigen with c-Src and beta-catenin". The Journal of Biological Chemistry. 276 (38): 35239–42. doi:10.1074/jbc.C100359200. PMID 11483589.
- ↑ Tang J, Feng GS, Li W (October 1997). "Induced direct binding of the adapter protein Nck to the GTPase-activating protein-associated protein p62 by epidermal growth factor". Oncogene. 15 (15): 1823–32. doi:10.1038/sj.onc.1201351. PMID 9362449.
- ↑ Li W, Hu P, Skolnik EY, Ullrich A, Schlessinger J (December 1992). "The SH2 and SH3 domain-containing Nck protein is oncogenic and a common target for phosphorylation by different surface receptors". Molecular and Cellular Biology. 12 (12): 5824–33. doi:10.1128/MCB.12.12.5824. PMC 360522. PMID 1333047.
- ↑ Chen M, She H, Davis EM, Spicer CM, Kim L, Ren R, Le Beau MM, Li W (September 1998). "Identification of Nck family genes, chromosomal localization, expression, and signaling specificity". The Journal of Biological Chemistry. 273 (39): 25171–8. doi:10.1074/jbc.273.39.25171. PMID 9737977.
- ↑ Tu Y, Li F, Wu C (December 1998). "Nck-2, a novel Src homology2/3-containing adaptor protein that interacts with the LIM-only protein PINCH and components of growth factor receptor kinase-signaling pathways". Molecular Biology of the Cell. 9 (12): 3367–82. doi:10.1091/mbc.9.12.3367. PMC 25640. PMID 9843575.
- ↑ Gauthier ML, Torretto C, Ly J, Francescutti V, O'Day DH (August 2003). "Protein kinase Calpha negatively regulates cell spreading and motility in MDA-MB-231 human breast cancer cells downstream of epidermal growth factor receptor". Biochemical and Biophysical Research Communications. 307 (4): 839–46. doi:10.1016/S0006-291X(03)01273-7. PMID 12878187.
- ↑ Bedrin MS, Abolafia CM, Thompson JF (July 1997). "Cytoskeletal association of epidermal growth factor receptor and associated signaling proteins is regulated by cell density in IEC-6 intestinal cells". Journal of Cellular Physiology. 172 (1): 126–36. doi:10.1002/(SICI)1097-4652(199707)172:1<126::AID-JCP14>3.0.CO;2-A. PMID 9207933.
- ↑ Sun J, Nanjundan M, Pike LJ, Wiedmer T, Sims PJ (May 2002). "Plasma membrane phospholipid scramblase 1 is enriched in lipid rafts and interacts with the epidermal growth factor receptor". Biochemistry. 41 (20): 6338–45. doi:10.1021/bi025610l. PMID 12009895.
- ↑ Sarmiento M, Puius YA, Vetter SW, Keng YF, Wu L, Zhao Y, Lawrence DS, Almo SC, Zhang ZY (July 2000). "Structural basis of plasticity in protein tyrosine phosphatase 1B substrate recognition". Biochemistry. 39 (28): 8171–9. doi:10.1021/bi000319w. PMID 10889023.
- ↑ Zhang ZY, Walsh AB, Wu L, McNamara DJ, Dobrusin EM, Miller WT (March 1996). "Determinants of substrate recognition in the protein-tyrosine phosphatase, PTP1". The Journal of Biological Chemistry. 271 (10): 5386–92. doi:10.1074/jbc.271.10.5386. PMID 8621392.
- ↑ 79.0 79.1 Tomic S, Greiser U, Lammers R, Kharitonenkov A, Imyanitov E, Ullrich A, Böhmer FD (September 1995). "Association of SH2 domain protein tyrosine phosphatases with the epidermal growth factor receptor in human tumor cells. Phosphatidic acid activates receptor dephosphorylation by PTP1C". The Journal of Biological Chemistry. 270 (36): 21277–84. doi:10.1074/jbc.270.36.21277. PMID 7673163.
- ↑ Keilhack H, Tenev T, Nyakatura E, Godovac-Zimmermann J, Nielsen L, Seedorf K, Böhmer FD (September 1998). "Phosphotyrosine 1173 mediates binding of the protein-tyrosine phosphatase SHP-1 to the epidermal growth factor receptor and attenuation of receptor signaling". The Journal of Biological Chemistry. 273 (38): 24839–46. doi:10.1074/jbc.273.38.24839. PMID 9733788.
- ↑ Wang SE, Wu FY, Shin I, Qu S, Arteaga CL (June 2005). "Transforming growth factor {beta} (TGF-{beta})-Smad target gene protein tyrosine phosphatase receptor type kappa is required for TGF-{beta} function". Molecular and Cellular Biology. 25 (11): 4703–15. doi:10.1128/MCB.25.11.4703-4715.2005. PMC 1140650. PMID 15899872.
- ↑ Lu Y, Brush J, Stewart TA (April 1999). "NSP1 defines a novel family of adaptor proteins linking integrin and tyrosine kinase receptors to the c-Jun N-terminal kinase/stress-activated protein kinase signaling pathway". The Journal of Biological Chemistry. 274 (15): 10047–52. doi:10.1074/jbc.274.15.10047. PMID 10187783.
- ↑ Soubeyran P, Kowanetz K, Szymkiewicz I, Langdon WY, Dikic I (March 2002). "Cbl-CIN85-endophilin complex mediates ligand-induced downregulation of EGF receptors". Nature. 416 (6877): 183–7. doi:10.1038/416183a. PMID 11894095.
- ↑ Szymkiewicz I, Kowanetz K, Soubeyran P, Dinarina A, Lipkowitz S, Dikic I (October 2002). "CIN85 participates in Cbl-b-mediated down-regulation of receptor tyrosine kinases". The Journal of Biological Chemistry. 277 (42): 39666–72. doi:10.1074/jbc.M205535200. PMID 12177062.
- ↑ Sakaguchi K, Okabayashi Y, Kido Y, Kimura S, Matsumura Y, Inushima K, Kasuga M (April 1998). "Shc phosphotyrosine-binding domain dominantly interacts with epidermal growth factor receptors and mediates Ras activation in intact cells". Molecular Endocrinology. 12 (4): 536–43. doi:10.1210/me.12.4.536. PMID 9544989.
- ↑ Qian X, Esteban L, Vass WC, Upadhyaya C, Papageorge AG, Yienger K, Ward JM, Lowy DR, Santos E (February 2000). "The Sos1 and Sos2 Ras-specific exchange factors: differences in placental expression and signaling properties". The EMBO Journal. 19 (4): 642–54. doi:10.1093/emboj/19.4.642. PMC 305602. PMID 10675333.
- ↑ Qian X, Vass WC, Papageorge AG, Anborgh PH, Lowy DR (February 1998). "N terminus of Sos1 Ras exchange factor: critical roles for the Dbl and pleckstrin homology domains". Molecular and Cellular Biology. 18 (2): 771–8. doi:10.1128/mcb.18.2.771. PMC 108788. PMID 9447973.
- ↑ Keely SJ, Calandrella SO, Barrett KE (April 2000). "Carbachol-stimulated transactivation of epidermal growth factor receptor and mitogen-activated protein kinase in T(84) cells is mediated by intracellular Ca2+, PYK-2, and p60(src)". The Journal of Biological Chemistry. 275 (17): 12619–25. doi:10.1074/jbc.275.17.12619. PMID 10777553.
- ↑ Sato K, Kimoto M, Kakumoto M, Horiuchi D, Iwasaki T, Tokmakov AA, Fukami Y (September 2000). "Adaptor protein Shc undergoes translocation and mediates up-regulation of the tyrosine kinase c-Src in EGF-stimulated A431 cells". Genes to Cells. 5 (9): 749–64. doi:10.1046/j.1365-2443.2000.00358.x. PMID 10971656.
- ↑ Xia L, Wang L, Chung AS, Ivanov SS, Ling MY, Dragoi AM, Platt A, Gilmer TM, Fu XY, Chin YE (August 2002). "Identification of both positive and negative domains within the epidermal growth factor receptor COOH-terminal region for signal transducer and activator of transcription (STAT) activation". The Journal of Biological Chemistry. 277 (34): 30716–23. doi:10.1074/jbc.M202823200. PMID 12070153.
- ↑ Yuan ZL, Guan YJ, Wang L, Wei W, Kane AB, Chin YE (November 2004). "Central role of the threonine residue within the p+1 loop of receptor tyrosine kinase in STAT3 constitutive phosphorylation in metastatic cancer cells". Molecular and Cellular Biology. 24 (21): 9390–400. doi:10.1128/MCB.24.21.9390-9400.2004. PMC 522220. PMID 15485908.
- ↑ Sehat B, Andersson S, Girnita L, Larsson O (July 2008). "Identification of c-Cbl as a new ligase for insulin-like growth factor-I receptor with distinct roles from Mdm2 in receptor ubiquitination and endocytosis". Cancer Research. 68 (14): 5669–77. doi:10.1158/0008-5472.CAN-07-6364. PMID 18632619.
- ↑ She HY, Rockow S, Tang J, Nishimura R, Skolnik EY, Chen M, Margolis B, Li W (September 1997). "Wiskott-Aldrich syndrome protein is associated with the adapter protein Grb2 and the epidermal growth factor receptor in living cells". Molecular Biology of the Cell. 8 (9): 1709–21. doi:10.1091/mbc.8.9.1709. PMC 305731. PMID 9307968.
- ↑ Shilo BZ (March 2003). "Signaling by the Drosophila epidermal growth factor receptor pathway during development". Experimental Cell Research. 284 (1): 140–9. doi:10.1016/S0014-4827(02)00094-0. PMID 12648473.
Further reading
- Carpenter G (1987). "Receptors for epidermal growth factor and other polypeptide mitogens". Annual Review of Biochemistry. 56 (1): 881–914. doi:10.1146/annurev.bi.56.070187.004313. PMID 3039909.
- Boonstra J, Rijken P, Humbel B, Cremers F, Verkleij A, van Bergen en Henegouwen P (May 1995). "The epidermal growth factor". Cell Biology International. 19 (5): 413–30. doi:10.1006/cbir.1995.1086. PMID 7640657.
- Carpenter G (August 2000). "The EGF receptor: a nexus for trafficking and signaling". BioEssays. 22 (8): 697–707. doi:10.1002/1521-1878(200008)22:8<697::AID-BIES3>3.0.CO;2-1. PMID 10918300.
- Filardo EJ (February 2002). "Epidermal growth factor receptor (EGFR) transactivation by estrogen via the G-protein-coupled receptor, GPR30: a novel signaling pathway with potential significance for breast cancer". The Journal of Steroid Biochemistry and Molecular Biology. 80 (2): 231–8. doi:10.1016/S0960-0760(01)00190-X. PMID 11897506.
- Tiganis T (January 2002). "Protein tyrosine phosphatases: dephosphorylating the epidermal growth factor receptor". IUBMB Life. 53 (1): 3–14. doi:10.1080/15216540210811. PMID 12018405.
- Di Fiore PP, Scita G (October 2002). "Eps8 in the midst of GTPases". The International Journal of Biochemistry & Cell Biology. 34 (10): 1178–83. doi:10.1016/S1357-2725(02)00064-X. PMID 12127568.
- Benaim G, Villalobo A (August 2002). "Phosphorylation of calmodulin. Functional implications". European Journal of Biochemistry / FEBS. 269 (15): 3619–31. doi:10.1046/j.1432-1033.2002.03038.x. PMID 12153558.
- Leu TH, Maa MC (January 2003). "Functional implication of the interaction between EGF receptor and c-Src". Frontiers in Bioscience. 8 (1–3): s28–38. doi:10.2741/980. PMID 12456372.
- Anderson NL, Anderson NG (November 2002). "The human plasma proteome: history, character, and diagnostic prospects". Molecular & Cellular Proteomics. 1 (11): 845–67. doi:10.1074/mcp.R200007-MCP200. PMID 12488461.
- Kari C, Chan TO, Rocha de Quadros M, Rodeck U (January 2003). "Targeting the epidermal growth factor receptor in cancer: apoptosis takes center stage". Cancer Research. 63 (1): 1–5. PMID 12517767.
- Bonaccorsi L, Muratori M, Carloni V, Zecchi S, Formigli L, Forti G, Baldi E (February 2003). "Androgen receptor and prostate cancer invasion". International Journal of Andrology. 26 (1): 21–5. doi:10.1046/j.1365-2605.2003.00375.x. PMID 12534934.
- Reiter J, Maihle NJ (May 2003). "Characterization and expression of novel 60-kDa and 110-kDa EGFR isoforms in human placenta". Annals of the New York Academy of Sciences. 995 (1): 39–47. doi:10.1111/j.1749-6632.2003.tb03208.x. PMID 12814937.
- Adams TE, McKern NM, Ward CW (June 2004). "Signalling by the type 1 insulin-like growth factor receptor: interplay with the epidermal growth factor receptor". Growth Factors. 22 (2): 89–95. doi:10.1080/08977190410001700998. PMID 15253384.
- Ferguson KM (November 2004). "Active and inactive conformations of the epidermal growth factor receptor". Biochemical Society Transactions. 32 (Pt 5): 742–5. doi:10.1042/BST0320742. PMID 15494003.
- Chao C, Hellmich MR (December 2004). "Bi-directional signaling between gastrointestinal peptide hormone receptors and epidermal growth factor receptor". Growth Factors. 22 (4): 261–8. doi:10.1080/08977190412331286900. PMID 15621729.
- Carlsson J, Ren ZP, Wester K, Sundberg AL, Heldin NE, Hesselager G, Persson M, Gedda L, Tolmachev V, Lundqvist H, Blomquist E, Nistér M (March 2006). "Planning for intracavitary anti-EGFR radionuclide therapy of gliomas. Literature review and data on EGFR expression". Journal of Neuro-Oncology. 77 (1): 33–45. doi:10.1007/s11060-005-7410-z. PMID 16200342.
- Scartozzi M, Pierantoni C, Berardi R, Antognoli S, Bearzi I, Cascinu S (April 2006). "Epidermal growth factor receptor: a promising therapeutic target for colorectal cancer". Analytical and Quantitative Cytology and Histology. 28 (2): 61–8. PMID 16637508.
- Prudkin L, Wistuba II (October 2006). "Epidermal growth factor receptor abnormalities in lung cancer. Pathogenetic and clinical implications". Annals of Diagnostic Pathology. 10 (5): 306–15. doi:10.1016/j.anndiagpath.2006.06.011. PMID 16979526.
- Ahmed SM, Salgia R (November 2006). "Epidermal growth factor receptor mutations and susceptibility to targeted therapy in lung cancer". Respirology. 11 (6): 687–92. doi:10.1111/j.1440-1843.2006.00887.x. PMID 17052295.
- Zhang X, Chang A (March 2007). "Somatic mutations of the epidermal growth factor receptor and non-small-cell lung cancer". Journal of Medical Genetics. 44 (3): 166–72. doi:10.1136/jmg.2006.046102. PMC 2598028. PMID 17158592.
- Cohenuram M, Saif MW (2007). "Epidermal growth factor receptor inhibition strategies in pancreatic cancer: past, present and the future". JOP. 8 (1): 4–15. PMID 17228128.
- Mellinghoff IK, Cloughesy TF, Mischel PS (January 2007). "PTEN-mediated resistance to epidermal growth factor receptor kinase inhibitors". Clinical Cancer Research. 13 (2 Pt 1): 378–81. doi:10.1158/1078-0432.CCR-06-1992. PMID 17255257.
- Nakamura JL (April 2007). "The epidermal growth factor receptor in malignant gliomas: pathogenesis and therapeutic implications". Expert Opinion on Therapeutic Targets. 11 (4): 463–72. doi:10.1517/14728222.11.4.463. PMID 17373877.
External links
- Epidermal Growth Factor Receptor at the US National Library of Medicine Medical Subject Headings (MeSH)
- Pages with broken file links
- Genes on human chromosome
- All articles with unsourced statements
- Articles with unsourced statements from October 2009
- Articles with invalid date parameter in template
- Articles with unsourced statements from July 2016
- Portal templates with all redlinked portals
- Tyrosine kinase receptors
- Oncogenes