Staphylococcus
| Staphylococcus | ||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|
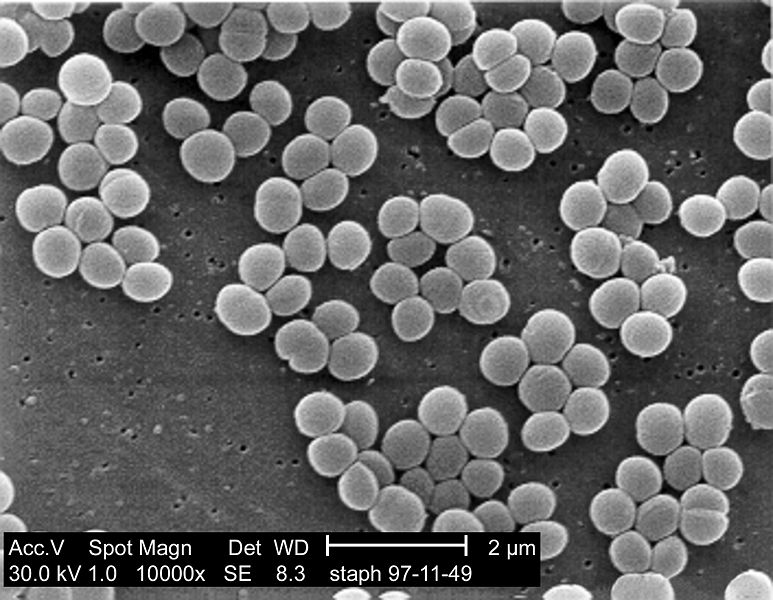 SEM micrograph of S. aureus colonies; note the grape-like clustering common to Staphylococcus species.
| ||||||||||||
| Scientific classification | ||||||||||||
| ||||||||||||
| Species | ||||||||||||
|
S. afermentans |
Editor-In-Chief: C. Michael Gibson, M.S., M.D. [2]
Overview
Staphylococcus (from the Template:Lang-el, staphylē, "grape" and κόκκος, kókkos, "granule") is a genus of Gram-positive bacteria. Under the microscope, they appear round (cocci), and form in grape-like clusters.[1]
The Staphylococcus genus includes at least 40 species. Of these, nine have two subspecies and one has three subspecies.[2] Most are harmless and reside normally on the skin and mucous membranes of humans and other organisms. Found worldwide, they are a small component of soil microbial flora.[3]
Taxonomy
The taxonomy is based on 16s rRNA sequences,[4] and most of the staphylococcal species fall into 11 clusters:
- S. aureus group – S. aureus, S. simiae
- S. auricularis group – S. auricularis
- S. carnosus group – S. carnosus, S. condimenti, S. massiliensis, S. piscifermentans, S. simulans
- S. epidermidis group – S. capitis, S. caprae, S. epidermidis, S. saccharolyticus
- S. haemolyticus group – S. devriesei, S. haemolyticus, S. hominis
- S. hyicus-intermedius group – S. chromogenes, S. felis, S. delphini, S. hyicus, S. intermedius, S. lutrae, S. microti, S. muscae, S. pseudintermedius, S. rostri, S. schleiferi
- S. lugdunensis group – S. lugdunensis
- S. saprophyticus group – S. arlettae, S. cohnii, S. equorum, S. gallinarum, S. kloosii, S. leei, S. nepalensis, S. saprophyticus, S. succinus, S. xylosus
- S. sciuri group – S. fleurettii, S. lentus, S. sciuri, S. stepanovicii, S. vitulinus
- S. simulans group – S. simulans
- S. warneri group – S. pasteuri, S. warneri
A twelfth group – that of S. caseolyticus – has now been moved to a new genus Macrococcus, the species of which are currently the closest known relatives of the Staphylococci.[5]
Subspecies
S. aureus subsp. aureus
S. aureus subsp. anaerobius
S. capitis subsp. capitis
S. capitis subsp. urealyticus
S. carnosus subsp. carnosus
S. carnosus subsp. utilis
S. cohnii subsp. cohnii
S. cohnii subsp. urealyticus
S. equorum subsp. equorum
S. equorum subsp. linens
S. hominis subsp. hominis
S. hominis subsp. novobiosepticus
S. saprophyticus subsp. bovis
S. saprophyticus subsp. saprophyticus
S. schleiferi subsp. coagulans
S. schleiferi subsp. schleiferi
S. sciuri subsp. carnaticus
S. sciuri subsp. rodentium
S. sciuri subsp. sciuri
S. succinus subsp. casei
S. succinus subsp. succinus
Notes
As with all generic names in binomial nomenclature, Staphylococcus is capitalized when used alone or with a specific species. Also, the abbreviations Staph and S. when used with a species (S. aureus) are correctly italicized and capitalized (though often errors in this are seen in popular literature). However, Staphylococcus is not capitalized or italicized when used in adjectival forms, as in a staphylococcal infection, or as the plural (staphylococci).[6]
The S. saprophyticus and S. sciuri groups are generally novobiocin-resistant, as is S. hominis subsp. novobiosepticus.
Members of the S. sciuri group are oxidase-positive due to their possession of the enzyme cytochrome c oxidase. This group is the only clade within the Staphylococci to possess this gene.
The S. sciuri group appears to be the closest relations to the genus Macrococcus.
Staphylococcus pulvereri has been shown to be a junior synonym of Staphylococcus vitulinus.[7]
Within these clades, the S. haemolyticus and S. simulans groups appear to be related, as do the S. aureus and S. epidermidis groups.[8]
S. lugdunensis appears to be related to the S. haemolyticus group.
S. croceolyticus may be related to S. haemolyticus, but this needs to be confirmed.
The taxonomic position of S. croceolyticus, S. leei, S. lyticans and S. pseudolugdunensis has yet to be clarified. The published descriptions of these species do not appear to have been validly published to date (2010).
Biochemical identification
Assignment of a strain to the genus Staphylococcus requires it to be a Gram-positive coccus that forms clusters, produces catalase, has an appropriate cell wall structure (including peptidoglycan type and teichoic acid presence) and G + C content of DNA in a range of 30–40 mol%.
Staphylococcus species can be differentiated from other aerobic and facultative anaerobic, Gram-positive cocci by several simple tests. Staphylococcus spp. are facultative anaerobes (capable of growth both aerobically and anaerobically). All species grow in the presence of bile salts.
It was believed that all species were coagulase-positive however it is now known that not all Staphylococcus are coagulase positive.[1][9][10]
Growth can also occur in a 6.5% NaCl solution. On Baird Parker medium, Staphylococcus spp. grow fermentatively, except for S. saprophyticus, which grows oxidatively. Staphylococcus spp. are resistant to bacitracin (0.04 U disc: resistance = < 10 mm zone of inhibition) and susceptible to furazolidone (100 μg disc: resistance = < 15 mm zone of inhibition). Further biochemical testing is needed to identify to the species level.
When the bacterium divides it divides along two axes, so forming clumps of bacteria. This is as opposed to streptococci which divide along one axis and so form chains (strep. meaning twisted or pliant).
Coagulase production
One of the most important phenotypical features used in the classification of staphylococci is their ability to produce coagulase, an enzyme that causes blood clot formation.
Six species are currently recognised as being coagulase-positive: S. aureus, S. delphini, S. hyicus, S. intermedius, S. lutrae,S. pseudintermedius and S. schleiferi subsp. coagulans. These species belong to two separate groups – the S. aureus (S. aureus alone) group and the S. hyicus-intermedius group (the remaining five). S. aureus can also be found as being coagulase-negative.
A seventh species has also been described – Staphylococcus leei – from patients with gastritis.[11]
S. aureus is coagulase-positive, meaning it produces coagulase. However, while the majority of S. aureus strains are coagulase-positive, some may be atypical in that they do not produce coagulase. S. aureus is catalase-positive (meaning that it can produce the enzyme catalase) and able to convert hydrogen peroxide (H2O2) to water and oxygen, which makes the catalase test useful to distinguish staphylococci from enterococci and streptococci.
S. pseudintermedius inhabits and sometimes infects the skin of domestic dogs and cats. This organism, too, can carry the genetic material that imparts multiple bacterial resistance. It is rarely implicated in infections in humans, as a zoonosis.
Coagulase-Negative Staphylococcus
S. epidermidis, a coagulase-negative species, is a commensal of the skin, but can cause severe infections in immune-suppressed patients and those with central venous catheters.
S. saprophyticus, another coagulase-negative species that is part of the normal vaginal flora, is predominantly implicated in genitourinary tract infections in sexually active young women.
In recent years, several other Staphylococcus species have been implicated in human infections, notably S. lugdunensis, S. schleiferi, and S. caprae.
Taxonomy of Coagulase-Negative Staphylococci Associated with Human Diseases
Genomics and molecular biology
The first S. aureus genomes to be sequenced were those of N315 and Mu50 in 2001. Many more complete S. aureus genomes have been submitted to the public databases, making it one of the most extensively sequenced bacteria. The use of genomic data is now widespread and provides a valuable resource for researchers working with S. aureus. Whole genome technologies, such as sequencing projects and microarrays, have shown an enormous variety of S. aureus strains. Each contains different combinations of surface proteins and different toxins. Relating this information to pathogenic behaviour is one of the major areas of staphylococcal research. The development of molecular typing methods has enabled the tracking of different strains of S. aureus. This may lead to better control of outbreak strains. A greater understanding of how the staphylococci evolve, especially due to the acquisition of mobile genetic elements encoding resistance and virulence genes is helping to identify new outbreak strains and may even prevent their emergence.[12]
The widespread incidence of antibiotic resistance across various strains of S. aureus, or across different species of Staphylococcus has been attributed to horizontal gene transfer of genes encoding antibiotic/metal resistance and virulence. A recent study demonstrated the extent of horizontal gene transfer among Staphylococcus to be much greater than previously expected, and encompasses genes with functions beyond antibiotic resistance and virulence, and beyond genes residing within the mobile genetic elements.[13]
Various strains of Staphylococcus are available from biological research centres, such as the National Collection of Type Cultures (NCTC).
Host range
Members of the genus Staphylococcus frequently colonize the skin and upper respiratory tracts of mammals and birds. Some species specificity has been observed in host range, such that the Staphylococcus species observed on some animals appear more rarely on more distantly related host species.[14] Some of the observed host specificity includes:
S. arlattae – chickens, goats
S. auricularis – deer, dogs, humans
S. capitis – humans
S. caprae – goats, humans
S. cohnii – chickens, humans
S. delphini – dolphins
S. devriesei – cattle
S. epidermiditis – humans
S. equorum – horses
S. felis – cats
S. fleurettii – goats
S. gallinarum – chickens, goats, pheasants
S. haemolyticus – humans, Cercocebus, Erythrocebus, Lemur, Macca, Microcebus, Pan
S. hyicus – pigs
S. leei – humans
S. lentus – goats, rabbits, sheep
S. lugdunensis – humans, goats
S. lutrae – otters
S. microti – voles (Microtus arvalis)
S. nepalensis – goats
S. pasteuri – humans, goats
S. pettenkoferi – humans
S. pseudintermedius – dogs
S. rostri – pigs
S. schleiferi – humans
S. sciuri – humans, dogs, goats
S. simiae – South American squirrel monkeys (Saimiri sciureus)
S. simulans – humans
S. warneri – humans, Cercopithecoidea, Pongidae
S. xylosus – humans
Clinical
Staphylococcus can cause a wide variety of diseases in humans and animals through either toxin production or penetration. Staphylococcal toxins are a common cause of food poisoning, as they can be produced by bacteria growing in improperly stored food items.
The most common sialadenitis is caused by staphylococci, as bacterial infections.[15]
See also
Notes and references
- ↑ 1.0 1.1 Ryan KJ, Ray CG, ed. (2004). Sherris Medical Microbiology (4th ed.). McGraw Hill. ISBN 0-8385-8529-9.
- ↑ Harris L.G, Foster S.J, Richards S. G. (2002). "An introduction to Staphylococcus aureus, and techniques for identifying and quantifying S. aureus adhesins in relation to adhesion to biomaterials: review" (PDF). European cells and materials. 4: 39–60. PMID 14562246.
- ↑ Madigan M, Martinko J, ed. (2005). Brock Biology of Microorganisms (11th ed.). Prentice Hall. ISBN 0-13-144329-1.
- ↑ Takahashi T, Satoh I, Kikuchi N. (1999). "Phylogenetic relationships of 38 taxa of the genus Staphylococcus based on 16S rRNA gene sequence analysis" (PDF). Int. J. Syst. Bacteriol. 49 (2): 725–728. doi:10.1099/00207713-49-2-725. PMID 10319495.
- ↑ Kloos WE, Ballard DN, George CG, Webster JA, Hubner RJ, Ludwig W, Schleifer KH, Fiedler F, Schubert K (1998). "Delimiting the genus Staphylococcus through description of Macrococcus caseolyticus gen. nov., comb. nov. and Macrococcus equipercicus sp. nov., and Macrococcus bovicus sp. nov. and Macrococcus carouselicus sp. nov" (PDF). Int J Syst Bacteriol. 48 (3): 859–877. doi:10.1099/00207713-48-3-859. PMID 9734040.
- ↑ See genus and species capitalization.
- ↑ Svec P., Vancanneyt M., Sedláek I., Engelbeen K., Stetina V., Swings, J. & Petrá, P. (2004). "Reclassification of Staphylococcus pulvereri Zakrzewska-Czerwiska et al. 1995 as a later synonym of Staphylococcus vitulinus Webster et al 1994" (PDF). Int. J. Syst. Evol. Microbiol. 54 (6): 2213–2215. doi:10.1099/ijs.0.63080-0. PMID 15545460.
- ↑ Ghebremedhin B, Layer F, König W, König B (2008). "Genetic classification and distinguishing of Staphylococcus species based on different partial gap, 16S rRNA, hsp60, rpoB, sodA, and tuf gene sequences". J. Clin. Microbiol. 46 (3): 1019–1025. doi:10.1128/JCM.02058-07. PMC 2268370. PMID 18174295.
- ↑ PreTest, Surgery, 12th ed., p. 88
- ↑ Matthews KR, Roberson J, Gillespie BE, Luther DA, Oliver SP (1997). "Identification and Differentiation of Coagulase-Negative Staphylococcus aureus by Polymerase Chain Reaction". Journal of Food Protection. 60 (6): 686–8.
- ↑ Jin M, Rosario W, Watler E, Calhoun DH (2004). "Development of a large-scale HPLC-based purification for the urease from Staphylococcus leei and determination of subunit structure" (PDF). Protein Expr. Purif. 34 (1): 111–117. doi:10.1016/j.pep.2003.10.01. PMID 14766306.
- ↑ Lindsay J (editor). (2008). Staphylococcus: Molecular Genetics. Caister Academic Press. ISBN 1-904455-29-8. [1].
- ↑ Chan CX, Beiko RG, Ragan MA (2011). "Lateral transfer of genes and gene fragments in Staphylococcus extends beyond mobile elements". J Bacteriol. 193 (15): 3964–3977. doi:10.1128/JB.01524-10. PMC 3147504. PMID 21622749.
- ↑ Kloos WE (1980). "Natural Populations of the Genus Staphylococcus". Annual Review of Microbiology. 34: 559–592. doi:10.1146/annurev.mi.34.100180.003015. PMID 7002032.
- ↑ "Sialoadenitis: inflammation of the salivary glands". The Medical Consumer's Advocate. 4 January 2001. Retrieved 2011-01-04.
External links
- Staphylococcus genomes and related information at PATRIC, a Bioinformatics Resource Center funded by NIAID