Notch signaling pathway

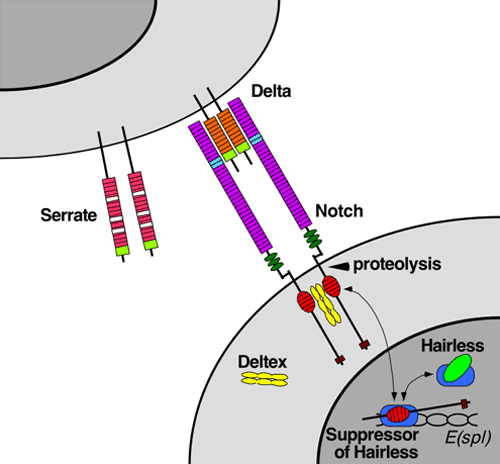

The Notch signaling pathway is a highly conserved cell signaling system present in most multicellular organisms. Notch is present in all metazoans, and vertebrates possess four different notch receptors, referred to as Notch1 to Notch4. The Notch receptor is a single-pass transmembrane receptor protein. It is a hetero-oligomer composed of a large extracellular portion, which associates in a calcium-dependent, non-covalent interaction with a smaller piece of the Notch protein composed of a short extracellular region, a single transmembrane-pass, and a small intracellular region.[1]
Discovery
The Notch gene was discovered in 1917 by Thomas Hunt Morgan, when it was first noticed in a strain of the fruit fly Drosophila melanogaster with notches apparent in their wingblades.[2][3] Its molecular analysis and sequencing was undertaken in the 1980s.[4][5]
Action
The Notch protein sits like a trigger spanning the cell membrane, with part of it inside and part outside. Ligand proteins binding to the extracellular domain induce proteolytic cleavage and release of the intracellular domain, which enters the cell nucleus to alter gene expression.[6]
Functions
The Notch signaling pathway is important for cell-cell communication, which involves gene regulation mechanisms that control multiple cell differentiation processes during embryonic and adult life. Notch signaling also has a role in the following processes:
- neuronal function and development[7][8]
- stabilization of arterial endothelial fate and angiogenesis[9]
- regulation of crucial cell communication events between endocardium and myocardium during both the formation of the valve primordial and ventricular development and differentiation [10]
- cardiac valve homeostasis, as well as implications in other human disorders involving the cardiovascular system[11]
- timely cell lineage specification of both endocrine and exocrine pancreas [12]
- influencing of binary fate decisions of cells that must choose between the secretory and absorptive lineages in the gut[13]
- expansion of the HSC compartment during bone development and participation in commitment to the osteoblastic lineage, suggesting a potential therapeutic role for Notch in bone regeneration and osteoporosis[14]
- regulation of cell-fate decision in mammary gland at several distinct development stages[15]
- possibly some non-nuclear mechanisms, such as control of the actin cytoskeleton through the tyrosine kinase Abl[16]
Notch signaling is dysregulated[1] in many cancers, and faulty Notch signaling is implicated in many diseases including T-ALL (T-cell acute lymphoblastic leukemia),[17] CADASIL (Cerebral Autosomal Dominant Arteriopathy with Sub-cortical Infarcts and Leukoencephalopathy), MS (Multiple Sclerosis), Tetralogy of Fallot, Alagille syndrome, and myriad other disease states.
Details of the pathway
Maturation of the Notch receptor involves cleavage at the prospective extracellular side during intracellular trafficking in the Golgi complex.[18] This results in a bipartite protein, composed of a large extracellular domain linked to the smaller transmembrane and intracellular domain. Binding of ligand promotes two proteolytic processing events; as a result of proteolysis, the intracellular domain is liberated and can enter the nucleus to engage other DNA-binding proteins and regulate gene expression.
Notch and most of its ligands are transmembrane proteins, so the cells expressing the ligands typically must be adjacent to the Notch expressing cell for signaling to occur. The Notch ligands are also single-pass transmembrane proteins and are members of the DSL (Delta/Serrate/LAG-2) family of proteins. In Drosophila melanogaster (the fruit fly), there are two ligands named Delta and Serrate. In mammals, the corresponding names are Delta-like and Jagged. In mammals there are multiple Delta-like and Jagged ligands, as well as possibly a variety of other ligands, such as F3/contactin[16].
In the nematode Caenorhabditis elegans, two genes encode homologous proteins, glp-1 and lin-12. There has been at least one report that suggests that some cells can send out processes that allow signaling to occur between cells that are as much as four or five cell diameters apart.
The Notch extracellular domain is composed primarily of small cysteine knot motifs called EGF-like repeats.[19] Notch 1, for example, has 36 of these repeats. Each EGF-like repeat is comprised of approximately 40 amino acids, and its structure is defined largely by six conserved cysteine residues that form three conserved disulfide bonds. Each EGF-like repeat can be modified by O-linked glycans at specific sites.[20] An O-glucose sugar may be added between the first and second conserved cysteines, and an O-fucose may be added between the second and third conserved cysteines. These sugars are added by an as-yet-unidentified O-glucosyltransferase, and GDP-fucose Protein O-fucosyltransferase 1 (POFUT1), respectively. The addition of O-fucose by POFUT1 is absolutely necessary for Notch function, and, without the enzyme to add O-fucose, all Notch proteins fail to function properly. As yet, the manner by which the glycosylation of Notch affects function is not completely understood.
The O-glucose on Notch can be further elongated to a trisaccharide with the addition of two xylose sugars by xylosyltransferases, and the O-fucose can be elongated to a tetrasaccharide by the ordered addition of an N-acetylglucosamine (GlcNAc) sugar by an N-Acetylglucosaminyltransferase called Fringe, the addition of a galactose by a galactosyltransferase, and the addition of a sialic acid by a sialyltransferase.[21]
To add another level of complexity, in mammals there are three Fringe GlcNAc-transferases, named Lunatic Fringe, Manic Fringe, and Radical Fringe. These enzymes are responsible for something called a "Fringe Effect" on Notch signaling.[22] If Fringe adds a GlcNAc to the O-fucose sugar, then the subsequent addition of a galactose and sialic acid will occur. In the presence of this tetrasaccharide, Notch signals strongly when it interacts with the Delta ligand, but has markedly inhibited signaling when interacting with the Jagged ligand.[23] The means by which this addition of sugar inhibits signaling through one ligand, and potentiates signaling through another is not clearly understood.
Once the Notch extracellular domain interacts with a ligand, an ADAM-family metalloprotease called TACE (Tumor Necrosis Factor Alpha Converting Enzyme) cleaves the Notch protein just outside the membrane.[24] This releases the extracellular portion of Notch, which continues to interact with the ligand. The ligand plus the Notch extracellular domain is then endocytosed by the ligand-expressing cell. There may be signaling effects in the ligand-expressing cell after endocytosis; this part of Notch signaling is a topic of active research. After this first cleavage, an enzyme called γ-secretase (which is implicated in Alzheimer's disease) cleaves the remaining part of the Notch protein just inside the inner leaflet of the cell membrane of the Notch-expressing cell. This releases the intracellular domain of the Notch protein, which then moves to the nucleus, where it can regulate gene expression by activating the transcription factor CSL.[16] Other proteins also participate in the intracellular portion of the Notch signaling cascade.
Triggering
Because most ligands are also transmembrane proteins, the receptor is normally triggered only from direct cell-to-cell contact. In this way, groups of cells can organise themselves, such that, if one cell expresses a given trait, this may be switched off in neighbour cells by the inter-cellular Notch signal. In this way, groups of cells influence one another to make large structures.
The Notch cascade consists of Notch and Notch ligands, as well as intracellular proteins transmitting the Notch signal to the cell's nucleus. The Notch/Lin-12/Glp-1 receptor family[25] was found to be involved in the specification of cell fates during development in Drosophila and C. elegans.[26] The Notch signaling pathway begins to inhibit new cell growth when adolescence is reached, and keeps neural networks stable in adulthood.
External links
- Diagram: Notch signaling pathway in Homo sapiens
- Diagram: Notch signaling in Drosophila
- Notch+Receptors at the US National Library of Medicine Medical Subject Headings (MeSH)
References
- ↑ Annika E. Walberg., (2002). "p300 and PCAF Act Cooperatively to Mediate Transcriptional Activation from Chromatin Templates by Notch Intracellular Domain In Vitro.". Molecular and Cellular Biology. 22 (22): 7812–7819. Unknown parameter
|month=ignored (help); Unknown parameter|coauthors=ignored (help); External link in|title=(help) - ↑ T. H. Morgan (1917). "The theory of the gene". The American Naturalist. 51 (609): 513–544. doi:10.1086/279629. Unknown parameter
|month=ignored (help) - ↑ Morgan, Thomas (1928). The theory of the gene (revised ed.). Yale University Press. pp. 77–81.
- ↑ Wharton KA (1985). "Nucleotide sequence from the neurogenic locus Notch implies a gene product that shares homology with proteins containing EGF-like repeats". Cell. 43 (3 pt 2): 567–581. doi:10.1016/0092-8674(85)90229-6. PMID 3935325. Unknown parameter
|coauthors=ignored (help); Unknown parameter|month=ignored (help) - ↑ Kidd S (1986). "Sequence of the Notch locus of Drosophila melanogaster: relationship of the encoded protein to mammalian clotting and growth factors". Mol Cell Biol. 6 (9): 3094–3108. PMID 3097517. Unknown parameter
|coauthors=ignored (help); Unknown parameter|month=ignored (help) - ↑ Franz Oswald (2001). "p300 Acts as a Transcriptional Coactivator for Mammalian Notch-1". Molecular and Cellular Biology. 21 (22): 7761–7774. doi:10.1128/MCB.21.22.7761-7774.2001. PMID 11604511. Unknown parameter
|coauthors=ignored (help); Unknown parameter|month=ignored (help); External link in|title=(help) - ↑ Gaiano N, (2002). "The role of notch in promoting glial and neural stem cell fates". Annual Reviews of Neuroscience. 25: 471. doi:10.1146/annurev.neuro.25.030702.130823. Text "PMID: 12052917
" ignored (help); Unknown parameter
|coauthors=ignored (help); Unknown parameter|month=ignored (help) - ↑ Bolos V, (2007). "Notch Signaling in Development and Cancer". Endocrine Reviews. 28: 339. doi:10.1210/er.2006-0046. PMID 17409286. Text "PMID: 17409286
" ignored (help); Unknown parameter
|coauthors=ignored (help); Unknown parameter|month=ignored (help) - ↑ Zhao-Jun Liu (2003). "Regulation of Notch1 and Dll4 by Vascular Endothelial Growth Factor in Arterial Endothelial Cells: Implications for Modulating Arteriogenesis and Angiogenesis". Molecular and Cellular Biology. 23 (1): 14–25. doi:10.1128/MCB.23.1.14-25.2003. PMID 12482957. Unknown parameter
|coauthors=ignored (help); Unknown parameter|month=ignored (help) - ↑ Joaquín Grego-Bessa; et al. (2007). "Notch Signaling Is Essential for Ventricular Chamber Development". Developmental Cell. 12 (3): 415–429. doi:10.1016/j.devcel.2006.12.011. Unknown parameter
|month=ignored (help) - ↑ The Notch signaling pathway in cardiac development and tissue homeostasis
- ↑ L. Charles Murtaugh (2003). "Notch signaling controls multiple steps of pancreatic differentiation". Proc Natl Acad Sci U S A. 100 (25): 14920–5. doi:10.1073/pnas.2436557100. Unknown parameter
|coauthors=ignored (help); Unknown parameter|month=ignored (help) - ↑ Guy R. Sander (2004). "Expression of Notch Receptors and Ligands in the Adult Gut". Journal of Histochemistry and Cytochemistry. 52 (4): 509–516. PMID 15034002. Unknown parameter
|coauthors=ignored (help) - ↑ Masuhiro Nobta; et al. (2005). "Critical Regulation of Bone Morphogenetic Protein-induced Osteoblastic Differentiation by Delta1/Jagged1-activated Notch1 Signaling". J. Biol. Chem. 280 (16): 15842–48. doi:10.1074/jbc.M412891200. PMID 15695512. Unknown parameter
|month=ignored (help) - ↑ Dontu, G. (2004). "Role of Notch signaling in cell-fate determination of human mammary stem/progenitor cells". Breast Cancer Res. 6 (2R605-15.): R605. doi:10.1186/bcr920. Unknown parameter
|coauthors=ignored (help) - ↑ 16.0 16.1 16.2 Eric C. Lai (2004). "Notch signaling: control of cell communication and cell fate.". Development. 131. Unknown parameter
|month=ignored (help); External link in|title=(help) - ↑ Sharma V.M., (2007). "The Notch1/c-Myc pathway in T cell leukemia.". Cell Cycle. 6 (8): 927–930. Text "PMID: 17404512
" ignored (help); Unknown parameter
|coauthors=ignored (help); Unknown parameter|month=ignored (help); External link in|title=(help) - ↑ Munro S (2000). "The notch signalling regulator fringe acts in the Golgi apparatus and requires the glycosyltransferase signature motif DXD". Curr Biol. 10 (14): 813–20. doi:10.1016/S0960-9822(00)00578-9. PMID 10899003. Unknown parameter
|month=ignored (help); Unknown parameter|coauthors=ignored (help) - ↑ Bing Ma, (2006). "Fucosylation in prokaryotes and eukaryotes.". Glycobiology. 16 (12). Unknown parameter
|coauthors=ignored (help); Unknown parameter|month=ignored (help); External link in|title=(help) - ↑ Shao L., (2002). "O-glycosylation of EGF repeats: identification and initial characterization of a UDP-glucose: protein O-glucosyltransferase.". Glycobiology. 12 (11): 763–770. Text "PMID: 12460944
" ignored (help); Unknown parameter
|coauthors=ignored (help); Unknown parameter|month=ignored (help); External link in|title=(help) - ↑ Lu L. (2006). "Roles of O-fucose glycans in notch signaling revealed by mutant mice.". Methods in Enzymology. 417: 127–136. doi:10.1016/S0076-6879(06)17010-X. Text "PMID: 17132502
" ignored (help); Unknown parameter
|coauthors=ignored (help); External link in|title=(help) - ↑ Thomas G.B., (2007). "The glycosyltransferase Fringe promotes Delta-Notch signaling between neurons and glia, and is required for subtype-specific glial gene expression.". Development. Unknown parameter
|coauthors=ignored (help); Text "PMID: 17215308 " ignored (help); Unknown parameter|month=ignored (help); External link in|title=(help) - ↑ LaVoie MJ., (2003). "The Notch ligands, Jagged and Delta, are sequentially processed by alpha-secretase and presenilin/gamma-secretase and release signaling fragments.". The Journal of Biological chemistry. 278 (38): 34427–37. Text "PMID: 12826675
" ignored (help); Unknown parameter
|coauthor=ignored (help); Unknown parameter|month=ignored (help); External link in|title=(help) - ↑ Brou C., (2000). "A novel proteolytic cleavage involved in Notch signaling: the role of the disintegrin-metalloprotease TACE". Molecular Cell. 5 (2): 207–16. doi:10.1016/S1097-2765(00)80417-7. Text "PMID: 10882063" ignored (help); Unknown parameter
|http://www.ncbi.nlm.nih.gov/entrez/query.fcgi?db=ignored (help); Unknown parameter|month=ignored (help); Unknown parameter|coauthors=ignored (help) - ↑ Artavanis-Tsakonas S, (1995). "Notch signaling.". Science. 268 (5208): 225–232. doi:10.1126/science.7716513. PMID 7716513. Text "PMID: 7716513
" ignored (help); Unknown parameter
|coauthors=ignored (help); Unknown parameter|month=ignored (help); External link in|title=(help) - ↑ Sinqson A., (1998). "The C. elegans spe-9 gene encodes a sperm transmembrane protein that contains EGF-like repeats and is required for fertilization.". Cell. 93 (1): 71–79. Text "PMID: 9546393
" ignored (help); Unknown parameter
|coauthors=ignored (help); Unknown parameter|month=ignored (help); External link in|title=(help)