Measles virus
| Measles | ||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|
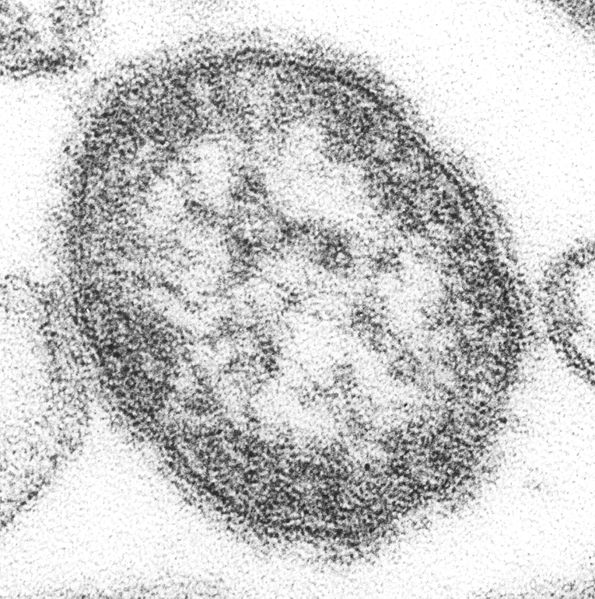 Measles virus electron micrograph
| ||||||||||||
| Virus classification | ||||||||||||
|
Editor-In-Chief: C. Michael Gibson, M.S., M.D. [1]; Associate Editor-In Chief: Joseph Nasr, M.D.[2]
|
Measles Microchapters |
|
Diagnosis |
|---|
|
Treatment |
|
Case Studies |
|
Measles virus On the Web |
|
American Roentgen Ray Society Images of Measles virus |
Overview
Measles virus (MeV) is a single-stranded, negative-sense, enveloped RNA virus of the genus Morbillivirus within the family Paramyxoviridae. Humans are the natural hosts of the virus; no animal reservoirs are known to exist.
Measles is highly contagious (R₀ ≈ 12–18) and spreads readily where vaccination coverage has fallen[1].
Persons with measles typically transmit the virus from ~4 days before to ~4 days after rash onset[1].
Microbiology & Clinical Correlates
- Syndrome. After an incubation of 10–14 days (range 7–23), illness begins with fever and the “3 Cs” (cough, coryza, conjunctivitis); Koplik spots may precede the maculopapular rash by 1–2 days[1].
- Systemic nature & complications. Because measles is systemic, complications (≈30% of cases) include diarrhea, pneumonia, otitis media, conjunctivitis; pneumonitis/giant-cell pneumonia occur mainly in immunosuppressed persons and young children[2].
- Encephalitis spectrum. Acute post-infectious encephalitis (days 0–7), measles-inclusion body encephalitis (1–6 months), and SSPE (years later) are rare but serious[3].
- Immune amnesia (clinical impact). Temporary B- and T-cell memory depletion increases susceptibility to secondary infections for months after recovery[4].
Additional properties: One antigenic type (epidemiologically); 100–200 nm; inactivated by heat, light, acidic pH, ether, trypsin; short environmental survival (<2 h); transmitted via respiratory secretions/aerosols.
Replication cycle
Entry
The measles virus has two envelope glycoproteins on the viral surface - hemagglutinin (H) and membrane fusion protein (F). These proteins are responsible for host cell binding and invasion. Three receptors for the H protein have been identified to date: complement regulatory molecule CD46, the signaling lymphocyte activation molecule (SLAM) and the cell adhesion molecule Nectin-4.[5]
- Entry receptors (H protein). CD46, SLAM/CD150[10], and nectin-4 are the three recognized receptors; CD150 (immune cells) and nectin-4 (epithelium) were identified in 2010–2011.
- Wild-type vs vaccine strains. Wild-type MeV primarily uses CD150[11], whereas vaccine strains mainly use CD46 (attenuation-associated shift).
- Cellular targeting in humans. CD150^hi memory T cells are preferential targets; CD150⁺ memory B cells and naïve B cells are also susceptible[12].
- Pathogenesis timeline: NEJM’s figure aligns incubation, contagious window (Day −4 to +4), MeV viremia, and immune-cell targets (CD150⁺ lymphocytes).
Replication cycle[1]
- Entry & fusion. H binds receptor (CD150 or nectin-4), F mediates membrane fusion → nucleocapsid release.
- Early spread & shedding. After initial respiratory/lymphoid infection, systemic viremia follows; patients are typically infectious Day −4 to +4 around rash appearance[1].
Immune modulation: immune amnesia & trained immunity
- Immune amnesia. During acute infection and for ≈5–12 months after resolution, measles can cause diminished pre-existing antibody repertoires and alter B-cell diversity, mechanistically explaining elevated post-measles morbidity/mortality from other infections.
- Vaccine does not cause amnesia; potential trained immunity. The appendix notes that measles vaccine strains do not cause adaptive or innate immune amnesia and summarizes evidence that MMR can induce “trained immunity” (innate immune reprogramming, including γδ T cells).
Evolution
The measles virus evolved from the formerly widespread rinderpest virus, which infects cattle.[13] Sequence analysis has suggested that the two viruses most probably diverged in the 11th and 12th centuries, though the periods as early as the 5th century fall within the 95% confidence interval of these calculations.[13]
Other analysis has suggested that the divergence may be even older because of the technique's tendency to underestimate ages when strong purifying selection is in action.[14] There is some linguistic evidence for an earlier origin within the seventh century.[15][16] The current epidemic strain evolved at the beginning of the 20th century—most probably between 1908 and 1943.[17]
Genotypes, antigenic stability, and vaccine cross-protection
Virology (brief). Measles virus (MV/MeV) is an enveloped, non-segmented, negative-sense RNA virus (Paramyxoviridae, Morbillivirus). It measures ~100–200 nm; two envelope glycoproteins drive entry: F (fusion) for membrane merger and H (hemagglutinin) for receptor binding. There is functionally one antigenic type in circulation; documented H changes have not translated into loss of vaccine effectiveness. MV is rapidly inactivated by heat, light, acidic pH, ether, and trypsin and survives <2 h in air/on surfaces.
Genotyping framework. For molecular surveillance, 24 MeV genotypes are recognized using the 450-bp N-gene window (“N-450”). Since 2018, genotypes B3, D4, D8, and H1 have been identified in global surveillance, with B3, D8, and H1 dominating 2024–2025 outbreaks[18].
(WHO defines 8 clades (A–H) with numbered subtypes (e.g., A1, D2); the genotyping scheme was introduced in 1998 and extended in 2002–2003; N-450 is the minimum sequence required for genotyping)
Antigenic stability (why vaccines still work). The surface glycoproteins H and F have largely retained their antigenic structure for decades; H-specific antibodies are major contributors to protection. A conserved immunodominant epitope on H overlaps the SLAM-binding site, so escape mutations tend to impair receptor binding/fitness rather than propagate[10][11][12].
Vaccine lineage & breadth. All currently used measles vaccines descend from genotype A (Edmonston lineage) and remain protective against circulating wild-type strains; the main text also reiterates that the vaccine is highly effective against all circulating genotypes[10][11][12].
Watch items (surveillance). Sub-genotype D4.2 has shown reduced binding to several monoclonal antibodies targeting major H epitopes; while clinical vaccine escape has not been shown, NEJM flags this as requiring close monitoring[19].
Elimination status notes. The predominant genotypes differ across regions and depend on whether endemic circulation persists. (indigenous transmission was interrupted in the United States and Australia by 2000 and in the Americas by 2002.)
References
- ↑ 1.0 1.1 1.2 1.3 1.4 Do, L.A.H. and Mulholland, K. (2025) “Measles 2025,” The New England journal of medicine [Preprint], (NEJMra2504516). Available at: https://doi.org/10.1056/NEJMra2504516.
- ↑ CDC (2025) Measles Symptoms and Complications, Measles (Rubeola). Available at: https://www.cdc.gov/measles/signs-symptoms/index.html (Accessed: September 22, 2025).
- ↑ Ferren, M., Horvat, B. and Mathieu, C. (2019) “Measles encephalitis: Towards new therapeutics,” Viruses, 11(11), p. 1017. Available at: https://doi.org/10.3390/v11111017.
- ↑ Mina, M.J. et al. (2015) “Long-term measles-induced immunomodulation increases overall childhood infectious disease mortality,” Science (New York, N.Y.), 348(6235), pp. 694–699. Available at: https://doi.org/10.1126/science.aaa3662.
- ↑ Lu G, Gao GF, Yan J (2013) The receptors and entry of measles virus: a review. Sheng Wu Gong Cheng Xue Bao 29(1):1–9
- ↑ Dorig RE, Marcil A, Chopra A, Richardson CD. The human CD46 molecule is a receptor for measles virus (Edmonston strain). Cell. 1993;75(2):295-305
- ↑ Naniche D, Varior-Krishnan G, Cervoni F, Wild TF, Rossi B, Rabourdin-Combe C, et al. Human membrane cofactor protein (CD46) acts as a cellular receptor for measles virus. J Virol. 1993;67(10):6025-32.
- ↑ Tatsuo H, Ono N, Tanaka K, Yanagi Y. SLAM (CDw150) is a cellular receptor for measles virus. Nature. 2000;406(6798):893-7.
- ↑ Muhlebach MD, Mateo M, Sinn PL, Prufer S, Uhlig KM, Leonard VH, et al. Adherens junction protein nectin-4 is the epithelial receptor for measles virus. Nature. 2011;480(7378):530-3.
- ↑ 10.0 10.1 10.2 Beaty SM, Lee B. Constraints on the Genetic and Antigenic Variability of Measles Virus. Viruses. 2016;8(4):109.
- ↑ 11.0 11.1 11.2 Munoz-Alia MA, Nace RA, Zhang L, Russell SJ. Serotypic evolution of measles virus is constrained by multiple co-dominant B cell epitopes on its surface glycoproteins. Cell Rep Med. 2021;2(4):100225.
- ↑ 12.0 12.1 12.2 Tahara M, Ohno S, Sakai K, Ito Y, Fukuhara H, Komase K, et al. The receptor-binding site of the measles virus hemagglutinin protein itself constitutes a conserved neutralizing epitope. J Virol. 2013;87(6):3583-6.
- ↑ 13.0 13.1 Furuse Y, Suzuki A, Oshitani H (2010). "Origin of measles virus: divergence from rinderpest virus between the 11th and 12th centuries". Virol. J. 7: 52. doi:10.1186/1743-422X-7-52. PMC 2838858. PMID 20202190.
- ↑ Template:Cite doi
- ↑ Griffin DE (2007). "Measles Virus". In Martin, Malcolm A.; Knipe, David M.; Fields, Bernard N.; Howley, Peter M.; Griffin, Diane; Lamb, Robert. Fields' virology (5th ed.). Philadelphia: Wolters Kluwer Health/Lippincott Williams & Wilkins. ISBN 0-7817-6060-7.
- ↑ McNeil, W. (1976). Plagues and Peoples. New York: Anchor Press/Doubleday. ISBN 0-385-11256-4.
- ↑ Pomeroy LW, Bjørnstad ON, Holmes EC (February 2008). "The evolutionary and epidemiological dynamics of the paramyxoviridae". J. Mol. Evol. 66 (2): 98–106. doi:10.1007/s00239-007-9040-x. PMC 3334863. PMID 18217182.
- ↑ CDC (2024) Genetic Analysis of Measles Viruses, Measles (Rubeola). Available at: https://www.cdc.gov/measles/php/laboratories/genetic-analysis.html (Accessed: September 22, 2025).
- ↑ Munoz-Alia MA, Muller CP, Russell SJ. Antigenic Drift Defines a New D4 Subgenotype of Measles Virus. J Virol. 2017;91(11).
index.php?title=Category:Pediatrics index.php?title=Category:Dermatology index.php?title=Category:Viral diseases index.php?title=Category:Mononegavirales index.php?title=Category:Ophthalmology index.php?title=Category:Otolaryngology index.php?title=Category:Pulmonology index.php?title=Category:Disease