Mycobacterium smegmatis: Difference between revisions
No edit summary |
|||
| Line 42: | Line 42: | ||
==Treatment== | ==Treatment== | ||
===Antimicrobial regimen=== | ===Antimicrobial regimen=== | ||
* | * [[Mycobacterium smegmatis]] <ref>{{Cite journal| doi = 10.1164/rccm.200604-571ST| issn = 1073-449X| volume = 175| issue = 4| pages = 367–416| last1 = Griffith| first1 = David E.| last2 = Aksamit| first2 = Timothy| last3 = Brown-Elliott| first3 = Barbara A.| last4 = Catanzaro| first4 = Antonino| last5 = Daley| first5 = Charles| last6 = Gordin| first6 = Fred| last7 = Holland| first7 = Steven M.| last8 = Horsburgh| first8 = Robert| last9 = Huitt| first9 = Gwen| last10 = Iademarco| first10 = Michael F.| last11 = Iseman| first11 = Michael| last12 = Olivier| first12 = Kenneth| last13 = Ruoss| first13 = Stephen| last14 = von Reyn| first14 = C. Fordham| last15 = Wallace| first15 = Richard J.| last16 = Winthrop| first16 = Kevin| last17 = ATS Mycobacterial Diseases Subcommittee| last18 = American Thoracic Society| last19 = Infectious Disease Society of America| title = An official ATS/IDSA statement: diagnosis, treatment, and prevention of nontuberculous mycobacterial diseases| journal = American Journal of Respiratory and Critical Care Medicine| date = 2007-02-15| pmid = 17277290}}</ref> | ||
:* | :* 1. '''Mild disease''' | ||
::* Preferred regimen: [[Doxycycline]] PO {{and}} [[ Trimethoprim sulfamethoxazole]] PO | |||
:* Preferred regimen: [[Doxycycline]] PO {{and}} [[ Trimethoprim sulfamethoxazole]] PO | :* 2. '''Severe disease''' | ||
::* Preferred regimen: [[Amikacin]] IV {{or}} [[Imipenem]] IV | |||
* Severe disease | |||
:* Preferred regimen: [[Amikacin]] IV {{or}} [[Imipenem]] IV | |||
==References== | ==References== | ||
Latest revision as of 16:02, 10 August 2015
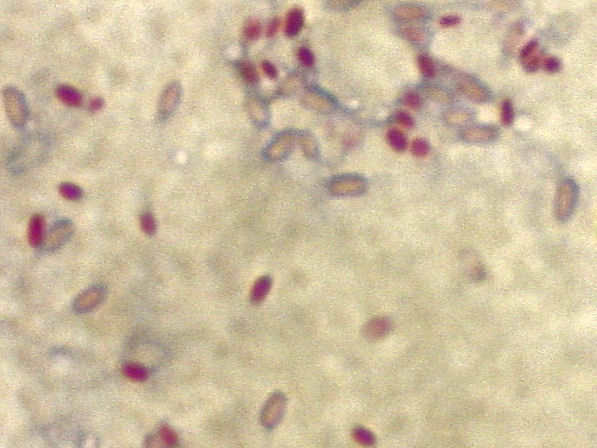
| style="background:#Template:Taxobox colour;"|Mycobacterium smegmatis | ||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|
 | ||||||||||||
| style="background:#Template:Taxobox colour;" | Scientific classification | ||||||||||||
| ||||||||||||
| Binomial name | ||||||||||||
| Mycobacterium smegmatis (Trevisan 1889) Lehmann & Neumann 1899 |
Editor-In-Chief: C. Michael Gibson, M.S., M.D. [1]
Overview
Mycobacterium smegmatis is an acid-fast bacterial species in the phylum Actinobacteria and the genus Mycobacterium. It is 3.0 to 5.0 µm long with a bacillus shape and can be stained by Ziehl-Neelsen method and the auramine-rhodamine fluorescent method. It was first reported in November 1884 by Lustgarten, who found a bacillus with the staining appearance of tubercle bacilli in syphilitic chancres. Subsequent to this, Alvarez and Tavel found organisms similar to that described by Lustgarten as well as in normal genital secretions (smegma). This organism was later named M. smegmatis.[1]
Pathogenicity
M. smegmatis is generally considered a non-pathogenic microorganism; however, in some very rare cases, it may cause disease.[2]
Use in research
M. smegmatis is useful for the research analysis of other Mycobacteria species in laboratory experiments. M. smegmatis is commonly used in work on the mycobacterium species due to its being a "fast grower" and non-pathogenic. M. smegmatis is a simple model that is easy to work with, i.e., with a fast doubling time and only requires a biosafety level 1 laboratory. The time and heavy infrastructure needed to work with pathogenic species prompted researchers to use M. smegmatis as a model for mycobacterial species. This species shares more than 2000 homologs with M. tuberculosis and shares the same unusual cell wall structure of M. tuberculosis and other mycobacterial species.[3] It is also capable of oxidizing carbon monoxide aerobically, as is M. tuberculosis.
The discovery of plasmids, phages, and mobile genetic elements has enabled the construction of dedicated gene-inactivation and gene reporter systems. The M. smegmatis mc2155 strain is hypertransformable, and is now the work-horse of mycobacterial genetics. Furthermore, it is readily cultivatable in most synthetic or complex laboratory media, where it can form visible colonies in 3–5 days. These properties make it a very attractive model organism for M. tuberculosis and other mycobacterial pathogens. M. smegmatis mc2155 is also used for the cultivation of mycobacteriophage. The complete genome of M. smegmatis has been sequenced by TIGR, and microarrays have been produced by PFGRC program (http://pfgrc.tigr.org/descriptionPages.shtml), adding further to its use as a model system to study mycobacteria.
Transformation
Transformation is a process by which a bacterial cell takes up DNA that had been released by another cell into the surrounding medium, and then incorporates that DNA into its own genome by homologous recombination (see Transformation (genetics)). Strains of M. smegmatis that have particularly efficient DNA repair machinery, as indicated by their greater resistance to the DNA damaging effects of agents such as UV and mitomycin C, proved to be the most capable of undergoing transformation.[4] This suggests that transformation in M. smegmatis is a DNA repair process, presumably a recombinational repair process, as it is in other bacterial species.[5]
Conjugation
Conjugal DNA transfer in M. smegmatis requires stable and extended contact between a donor and a recipient strain, is DNase resistant, and the transferred DNA is incorporated into the recipient’s chromosome by homologous recombination. However, in contrast to the well-known E. coli Hfr conjugation system, in M. smegmatis all regions of the chromosome are transferred with comparable efficiencies and mycobacterial conjugation is chromosome, rather than plasmid based. Gray et al.[6] reported substantial blending of the parental genomes resulting from conjugation and referred to this blending as reminiscent of that seen in the meiotic products of sexual reproduction (see Origin of sexual reproduction)
DNA repair
M. smegmatis relies on DNA repair pathways to resist DNA damage. Double-strand breaks are especially threatening to bacterial viability. M. smegmatis has three options for repairing double-strand breaks; homologous recombination (HR), non-homologous end joining (NHEJ), and single-strand annealing (SSA).[7] The HR pathway of M. smegmatis is the major determinant of resistance to ionizing radiation and oxidative DNA damage. This pathway involves exchange of information between a damaged chromosome and another homologous chromosome in the same cell. It depends on the RecA protein that catalyzes strand exchange and the Adn protein that acts as a presynaptic nuclease.[7] HR is an accurate repair process and is the preferred pathway during logarithmic growth.[8]
The NHEJ pathway for repairing double-strand breaks involves the rejoining of the broken ends. It does not depend on a second homologous chromosome. This pathway requires the Ku protein and a specialized poly-functional ATP-dependent DNA ligase (ligase D).[9] NHEJ is efficient but inaccurate. Sealing of blunt DNA ends within a functional gene sequence occurs with a mutation frequency of about 50%.[9] NHEJ is the preferred pathway during stationary phase, and it protects M. smegmatis against the harmful effects of desiccation.[8]
SSA is employed as a repair pathway when a double-strand break arises between direct repeat sequences in DNA. SSA involves single-strand resection, annealing of the repeats, flap removal, gap filling and ligation. In M. smegmatis the SSA pathway depends on the RecBCD helicase-nuclease.[7]
Treatment
Antimicrobial regimen
- 1. Mild disease
- Preferred regimen: Doxycycline PO AND Trimethoprim sulfamethoxazole PO
- 2. Severe disease
References
- ↑ GORDON, RE; SMITH, MM (July 1953). "Rapidly growing, acid fast bacteria. I. Species' descriptions of Mycobacterium phlei Lehmann and Neumann and Mycobacterium smegmatis (Trevisan) Lehmann and Neumann". Journal of bacteriology. 66 (1): 41–8. PMC 357089. PMID 13069464.
- ↑ Reyrat, Jean-Marc; Kahn, Daniel (1 October 2001). "Mycobacterium smegmatis: an absurd model for tuberculosis?". Trends in Microbiology. 9 (10): 472–473. doi:10.1016/S0966-842X(01)02168-0.
- ↑ King, Gary (2003). "Uptake of carbon monoxide and hydrogen at environmentally relevant concentrations by mycobacteria". Applied and Environmental Microbiology. 69: 7266–7272. doi:10.1128/aem.69.12.7266-7272.2003.
- ↑ Norgard MV, Imaeda T (1978). "Physiological factors involved in the transformation of Mycobacterium smegmatis". J. Bacteriol. 133 (3): 1254–62. PMC 222159. PMID 641008.
- ↑ Michod RE, Bernstein H, Nedelcu AM (2008). "Adaptive value of sex in microbial pathogens". Infect. Genet. Evol. 8 (3): 267–85. doi:10.1016/j.meegid.2008.01.002. PMID 18295550.
- ↑ Gray TA, Krywy JA, Harold J, Palumbo MJ, Derbyshire KM (2013). "Distributive conjugal transfer in mycobacteria generates progeny with meiotic-like genome-wide mosaicism, allowing mapping of a mating identity locus". PLoS Biol. 11 (7): e1001602. doi:10.1371/journal.pbio.1001602. PMC 3706393. PMID 23874149.
- ↑ 7.0 7.1 7.2 Gupta R, Barkan D, Redelman-Sidi G, Shuman S, Glickman MS (2011). "Mycobacteria exploit three genetically distinct DNA double-strand break repair pathways". Mol. Microbiol. 79 (2): 316–30. doi:10.1111/j.1365-2958.2010.07463.x. PMC 3812669. PMID 21219454.
- ↑ 8.0 8.1 Pitcher RS, Green AJ, Brzostek A, Korycka-Machala M, Dziadek J, Doherty AJ (2007). "NHEJ protects mycobacteria in stationary phase against the harmful effects of desiccation". DNA Repair (Amst.). 6 (9): 1271–6. doi:10.1016/j.dnarep.2007.02.009. PMID 17360246.
- ↑ 9.0 9.1 Gong C, Bongiorno P, Martins A, Stephanou NC, Zhu H, Shuman S, Glickman MS (2005). "Mechanism of nonhomologous end-joining in mycobacteria: a low-fidelity repair system driven by Ku, ligase D and ligase C". Nat. Struct. Mol. Biol. 12 (4): 304–12. doi:10.1038/nsmb915. PMID 15778718.
- ↑ Griffith, David E.; Aksamit, Timothy; Brown-Elliott, Barbara A.; Catanzaro, Antonino; Daley, Charles; Gordin, Fred; Holland, Steven M.; Horsburgh, Robert; Huitt, Gwen; Iademarco, Michael F.; Iseman, Michael; Olivier, Kenneth; Ruoss, Stephen; von Reyn, C. Fordham; Wallace, Richard J.; Winthrop, Kevin; ATS Mycobacterial Diseases Subcommittee; American Thoracic Society; Infectious Disease Society of America (2007-02-15). "An official ATS/IDSA statement: diagnosis, treatment, and prevention of nontuberculous mycobacterial diseases". American Journal of Respiratory and Critical Care Medicine. 175 (4): 367–416. doi:10.1164/rccm.200604-571ST. ISSN 1073-449X. PMID 17277290.